| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2021-10-13 04:27:42 UTC |
|---|
| HMDB ID | HMDB0000194 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
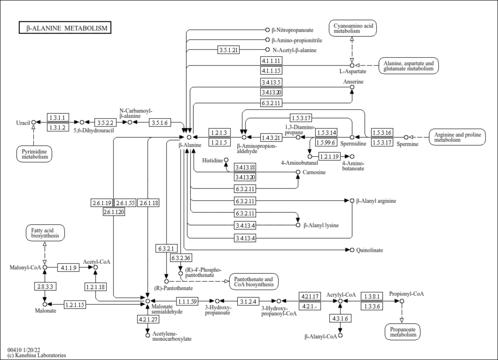
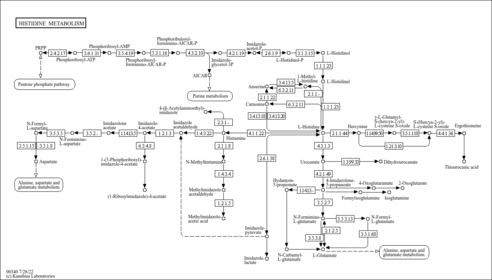
| Common Name | Anserine |
|---|
| Description | Anserine (beta-alanyl-N-3-methylhistidine) is a dipeptide containing beta-alanine and 3-methylhistidine. It is a derivative of carnosine, which had been methylated. The methyl group of anserine is added to carnosine by the enzyme S-adenosylmethionine: carnosine N-methyltransferase (PMID: 29484990 ). The enzyme is closely related to histamine N-methyltransferase and appears to be present in a majority of anserine-producing species (PMID: 23705015 ). Anserine is a generally a more metabolically stable derivative of carnosine. Anserine can be found in the skeletal muscle and brain of certain mammals (rabbits, cattle), migratory fish and birds. This dipeptide is normally absent from human tissues and body fluids, and its appearance there is usually an artifact of diet. Anserine can also arise from serum carnosinase deficiency. (OMIM 212200 ). Anserine was first discovered in goose muscle in 1929, and was named after this extraction (anser is Latin for goose). Anserine, which is water-soluble, is found at high levels in the muscles of different non-human vertebrates, with poultry, rabbit, tuna, plaice, and salmon having generally higher contents than other marine foods, beef, or pork (PMID: 31908682 ). An increase of urinary anserine excretion has been found in humans after the consumption of chicken, rabbit, and tuna and has been associated with intake of chicken, salmon, and, to a lesser extent, beef (PMID: 31908682 ). Anserine can undergo cleavage to give rise to 3-methylhistidine.(3-MH). The dipeptide balenine, common in some whales, cleaves to form 1-methylhistidine (1-MH) (PMID: 31908682 ). There is considerable confusion with regard to the nomenclature of the methylated nitrogen atoms on the imidazole ring of histidine and other histidine-containing peptides such as anserine. In particular, older literature (mostly prior to the year 2000) designated anserine (N-pi methylated) as beta-alanyl-N1-methyl-histidine, whereas according to standard IUPAC nomenclature, anserine is correctly named as beta-alanyl-N3-methyl-histidine. As a result, many papers published prior to the year 2000 incorrectly identified 1MH as a specific marker for dietary consumption of certain foods or various pathophysiological effects when they really were referring to 3MH or vice versa (PMID: 24137022 ). In particular balenine (a whale or snake-specific dipeptide with 1MH) was often confused with anserine (the poultry dipeptide with 3MH). An animal model study of Alzheimer's disease using mice found that treatment with anserine reduced memory loss (PMID: 28974740 ). Anserine reduced glial inflammatory activity (particularly of astrocyte). The study also found that anserine-treated mice had greater pericyte surface area. The greater area of pericytes was commensurate with improved memory. The anserine-treated mice overall performed better on a spatial memory test (Morris Water Maze) (PMID: 28974740 ). A human study on 84 elderly subjects showed that subjects who took anserine and carnosine supplements for one year showed increased blood flow in the prefrontal cortex on MRI (PMID: 29896423 ). |
|---|
| Structure | CN1C=NC=C1C[C@H](NC(=O)CCN)C(O)=O InChI=1S/C10H16N4O3/c1-14-6-12-5-7(14)4-8(10(16)17)13-9(15)2-3-11/h5-6,8H,2-4,11H2,1H3,(H,13,15)(H,16,17)/t8-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| beta-Alanyl-3-methyl-L-histidine | ChEBI | | beta-Alanyl-N(pai)-methyl-L-histidine | ChEBI | | b-Alanyl-3-methyl-L-histidine | Generator | | Β-alanyl-3-methyl-L-histidine | Generator | | b-Alanyl-N(pai)-methyl-L-histidine | Generator | | Β-alanyl-N(pai)-methyl-L-histidine | Generator | | L-Anserine | HMDB | | L-N-b-Alanyl-3-methyl-histidine | HMDB | | L-N-beta-Alanyl-3-methyl-histidine | HMDB | | N-b-Alanyl-3-methyl-L-histidine | HMDB | | N-beta-Alanyl-3-methyl-L-histidine | HMDB | | Balanine | HMDB | | Beta Alanyl 3 methylhistidine | HMDB | | Beta-Alanyl-3-methylhistidine | HMDB | | Ophidine | HMDB |
|
|---|
| Chemical Formula | C10H16N4O3 |
|---|
| Average Molecular Weight | 240.259 |
|---|
| Monoisotopic Molecular Weight | 240.122240398 |
|---|
| IUPAC Name | (2S)-2-(3-aminopropanamido)-3-(1-methyl-1H-imidazol-5-yl)propanoic acid |
|---|
| Traditional Name | anserine |
|---|
| CAS Registry Number | 584-85-0 |
|---|
| SMILES | CN1C=NC=C1C[C@H](NC(=O)CCN)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C10H16N4O3/c1-14-6-12-5-7(14)4-8(10(16)17)13-9(15)2-3-11/h5-6,8H,2-4,11H2,1H3,(H,13,15)(H,16,17)/t8-/m0/s1 |
|---|
| InChI Key | MYYIAHXIVFADCU-QMMMGPOBSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as hybrid peptides. Hybrid peptides are compounds containing at least two different types of amino acids (alpha, beta, gamma, delta) linked to each other through a peptide bond. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Peptidomimetics |
|---|
| Sub Class | Hybrid peptides |
|---|
| Direct Parent | Hybrid peptides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Hybrid peptide
- Histidine or derivatives
- N-acyl-alpha-amino acid
- N-acyl-alpha amino acid or derivatives
- Alpha-amino acid or derivatives
- Imidazolyl carboxylic acid derivative
- N-substituted imidazole
- Azole
- Imidazole
- Heteroaromatic compound
- Amino acid or derivatives
- Amino acid
- Carboximidic acid
- Carboximidic acid derivative
- Carboxylic acid derivative
- Carboxylic acid
- Azacycle
- Monocarboxylic acid or derivatives
- Organoheterocyclic compound
- Organic 1,3-dipolar compound
- Propargyl-type 1,3-dipolar organic compound
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Primary aliphatic amine
- Carbonyl group
- Organic nitrogen compound
- Amine
- Organic oxygen compound
- Organonitrogen compound
- Organooxygen compound
- Primary amine
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | |
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 1.6 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 9.0563 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 8.8 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 422.6 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 349.7 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 243.5 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 50.8 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 162.7 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 61.3 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 299.7 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 242.0 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 1006.0 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 541.9 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 45.1 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 626.8 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 186.2 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 257.8 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 742.2 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 746.7 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 392.7 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Anserine,1TMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN)C(=O)O[Si](C)(C)C | 2402.9 | Semi standard non polar | 33892256 | | Anserine,1TMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C)C(=O)O | 2498.8 | Semi standard non polar | 33892256 | | Anserine,1TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN)[Si](C)(C)C | 2320.9 | Semi standard non polar | 33892256 | | Anserine,2TMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C)C(=O)O[Si](C)(C)C | 2514.8 | Semi standard non polar | 33892256 | | Anserine,2TMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C)C(=O)O[Si](C)(C)C | 2352.8 | Standard non polar | 33892256 | | Anserine,2TMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C)C(=O)O[Si](C)(C)C | 3248.9 | Standard polar | 33892256 | | Anserine,2TMS,isomer #2 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN)[Si](C)(C)C | 2326.9 | Semi standard non polar | 33892256 | | Anserine,2TMS,isomer #2 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN)[Si](C)(C)C | 2276.0 | Standard non polar | 33892256 | | Anserine,2TMS,isomer #2 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN)[Si](C)(C)C | 3439.2 | Standard polar | 33892256 | | Anserine,2TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN[Si](C)(C)C)[Si](C)(C)C | 2403.0 | Semi standard non polar | 33892256 | | Anserine,2TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN[Si](C)(C)C)[Si](C)(C)C | 2415.1 | Standard non polar | 33892256 | | Anserine,2TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN[Si](C)(C)C)[Si](C)(C)C | 3309.7 | Standard polar | 33892256 | | Anserine,2TMS,isomer #4 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C)[Si](C)(C)C)C(=O)O | 2626.2 | Semi standard non polar | 33892256 | | Anserine,2TMS,isomer #4 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C)[Si](C)(C)C)C(=O)O | 2523.0 | Standard non polar | 33892256 | | Anserine,2TMS,isomer #4 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C)[Si](C)(C)C)C(=O)O | 3481.3 | Standard polar | 33892256 | | Anserine,3TMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN[Si](C)(C)C)[Si](C)(C)C | 2413.9 | Semi standard non polar | 33892256 | | Anserine,3TMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN[Si](C)(C)C)[Si](C)(C)C | 2405.7 | Standard non polar | 33892256 | | Anserine,3TMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN[Si](C)(C)C)[Si](C)(C)C | 2927.8 | Standard polar | 33892256 | | Anserine,3TMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C)[Si](C)(C)C)C(=O)O[Si](C)(C)C | 2622.7 | Semi standard non polar | 33892256 | | Anserine,3TMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C)[Si](C)(C)C)C(=O)O[Si](C)(C)C | 2471.8 | Standard non polar | 33892256 | | Anserine,3TMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C)[Si](C)(C)C)C(=O)O[Si](C)(C)C | 3053.3 | Standard polar | 33892256 | | Anserine,3TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN([Si](C)(C)C)[Si](C)(C)C)[Si](C)(C)C | 2558.7 | Semi standard non polar | 33892256 | | Anserine,3TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN([Si](C)(C)C)[Si](C)(C)C)[Si](C)(C)C | 2563.4 | Standard non polar | 33892256 | | Anserine,3TMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN([Si](C)(C)C)[Si](C)(C)C)[Si](C)(C)C | 3114.6 | Standard polar | 33892256 | | Anserine,4TMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN([Si](C)(C)C)[Si](C)(C)C)[Si](C)(C)C | 2582.3 | Semi standard non polar | 33892256 | | Anserine,4TMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN([Si](C)(C)C)[Si](C)(C)C)[Si](C)(C)C | 2540.2 | Standard non polar | 33892256 | | Anserine,4TMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C)N(C(=O)CCN([Si](C)(C)C)[Si](C)(C)C)[Si](C)(C)C | 2792.4 | Standard polar | 33892256 | | Anserine,1TBDMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN)C(=O)O[Si](C)(C)C(C)(C)C | 2676.5 | Semi standard non polar | 33892256 | | Anserine,1TBDMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C(C)(C)C)C(=O)O | 2732.7 | Semi standard non polar | 33892256 | | Anserine,1TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN)[Si](C)(C)C(C)(C)C | 2598.1 | Semi standard non polar | 33892256 | | Anserine,2TBDMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2949.3 | Semi standard non polar | 33892256 | | Anserine,2TBDMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2750.8 | Standard non polar | 33892256 | | Anserine,2TBDMS,isomer #1 | CN1C=NC=C1C[C@H](NC(=O)CCN[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 3277.2 | Standard polar | 33892256 | | Anserine,2TBDMS,isomer #2 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN)[Si](C)(C)C(C)(C)C | 2812.6 | Semi standard non polar | 33892256 | | Anserine,2TBDMS,isomer #2 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN)[Si](C)(C)C(C)(C)C | 2704.1 | Standard non polar | 33892256 | | Anserine,2TBDMS,isomer #2 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN)[Si](C)(C)C(C)(C)C | 3410.3 | Standard polar | 33892256 | | Anserine,2TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2915.2 | Semi standard non polar | 33892256 | | Anserine,2TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2794.2 | Standard non polar | 33892256 | | Anserine,2TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3309.7 | Standard polar | 33892256 | | Anserine,2TBDMS,isomer #4 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O | 3066.3 | Semi standard non polar | 33892256 | | Anserine,2TBDMS,isomer #4 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O | 2882.3 | Standard non polar | 33892256 | | Anserine,2TBDMS,isomer #4 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O | 3461.2 | Standard polar | 33892256 | | Anserine,3TBDMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3090.0 | Semi standard non polar | 33892256 | | Anserine,3TBDMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2969.7 | Standard non polar | 33892256 | | Anserine,3TBDMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3131.5 | Standard polar | 33892256 | | Anserine,3TBDMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 3292.1 | Semi standard non polar | 33892256 | | Anserine,3TBDMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 3057.7 | Standard non polar | 33892256 | | Anserine,3TBDMS,isomer #2 | CN1C=NC=C1C[C@H](NC(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 3202.5 | Standard polar | 33892256 | | Anserine,3TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3240.2 | Semi standard non polar | 33892256 | | Anserine,3TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3100.4 | Standard non polar | 33892256 | | Anserine,3TBDMS,isomer #3 | CN1C=NC=C1C[C@@H](C(=O)O)N(C(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3244.9 | Standard polar | 33892256 | | Anserine,4TBDMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3421.1 | Semi standard non polar | 33892256 | | Anserine,4TBDMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3240.6 | Standard non polar | 33892256 | | Anserine,4TBDMS,isomer #1 | CN1C=NC=C1C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N(C(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3106.4 | Standard polar | 33892256 |
|
|---|
| Disease References | | Alzheimer's disease |
|---|
- Fonteh AN, Harrington RJ, Tsai A, Liao P, Harrington MG: Free amino acid and dipeptide changes in the body fluids from Alzheimer's disease subjects. Amino Acids. 2007 Feb;32(2):213-24. Epub 2006 Oct 10. [PubMed:17031479 ]
| | Colorectal cancer |
|---|
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| General References | - Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Harris RC, Tallon MJ, Dunnett M, Boobis L, Coakley J, Kim HJ, Fallowfield JL, Hill CA, Sale C, Wise JA: The absorption of orally supplied beta-alanine and its effect on muscle carnosine synthesis in human vastus lateralis. Amino Acids. 2006 May;30(3):279-89. Epub 2006 Mar 24. [PubMed:16554972 ]
- Pagnano MW, Hanssen AD, Lewallen DG, Stuart MJ: Flexion instability after primary posterior cruciate retaining total knee arthroplasty. Clin Orthop Relat Res. 1998 Nov;(356):39-46. [PubMed:9917666 ]
- Abe H, Okuma E, Sekine H, Maeda A, Yoshiue S: Human urinary excretion of L-histidine-related compounds after ingestion of several meats and fish muscle. Int J Biochem. 1993 Sep;25(9):1245-9. [PubMed:8224369 ]
- Kang I, Han SW: Anserine bursitis in patients with osteoarthritis of the knee. South Med J. 2000 Feb;93(2):207-9. [PubMed:10701790 ]
- Magana Loarte JE, Perez Franco J, Sanchez Sanchez G: [Is therapy with local infiltrations feasible in primary care consultations?]. Aten Primaria. 1999 Jan;23(1):4-7. [PubMed:10079554 ]
- Tan KM, Candlish JK: Carnosine and anserine as modulators of neutrophil function. Clin Lab Haematol. 1998 Aug;20(4):239-44. [PubMed:9777271 ]
- Boldyrev A, Abe H: Metabolic transformation of neuropeptide carnosine modifies its biological activity. Cell Mol Neurobiol. 1999 Feb;19(1):163-75. [PubMed:10079975 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
- Kwiatkowski S, Kiersztan A, Drozak J: Biosynthesis of Carnosine and Related Dipeptides in Vertebrates. Curr Protein Pept Sci. 2018;19(8):771-789. doi: 10.2174/1389203719666180226155657. [PubMed:29484990 ]
- Drozak J, Chrobok L, Poleszak O, Jagielski AK, Derlacz R: Molecular identification of carnosine N-methyltransferase as chicken histamine N-methyltransferase-like protein (hnmt-like). PLoS One. 2013 May 21;8(5):e64805. doi: 10.1371/journal.pone.0064805. Print 2013. [PubMed:23705015 ]
|
|---|