| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Expected but not Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2022-03-07 02:49:08 UTC |
|---|
| HMDB ID | HMDB0001052 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | (S)-3-Hydroxyisobutyryl-CoA |
|---|
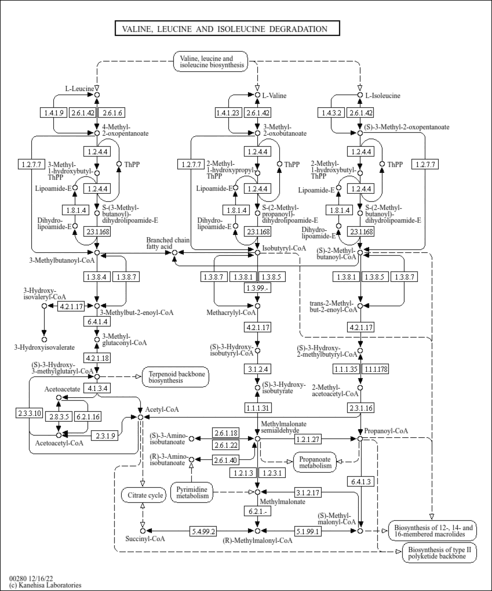
| Description | (S)-3-Hydroxyisobutyryl-CoA is s metabolite of 3-hydroxyisobutyryl-CoA hydrolase (EC 3.1.2.4 ) during beta-alanine metabolism (KEGG 00410), propanoate metabolism (KEGG 00640), and valine, leucine and isoleucine degradation (KEGG 00280). Deficiencies of this enzyme in valine degradation can result in hypotonia, poor feeding, motor delay, and subsequent neurological regression in infancy, episodes of ketoacidosis and Leigh-like changes in the basal ganglia on a magnetic resonance imaging scan (PMID 17160907 ). |
|---|
| Structure | CC(CO)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OCC1OC(C(O)C1OP(O)(O)=O)N1C=NC2=C(N)N=CN=C12 InChI=1S/C25H42N7O18P3S/c1-13(8-33)24(38)54-7-6-27-15(34)4-5-28-22(37)19(36)25(2,3)10-47-53(44,45)50-52(42,43)46-9-14-18(49-51(39,40)41)17(35)23(48-14)32-12-31-16-20(26)29-11-30-21(16)32/h11-14,17-19,23,33,35-36H,4-10H2,1-3H3,(H,27,34)(H,28,37)(H,42,43)(H,44,45)(H2,26,29,30)(H2,39,40,41) |
|---|
| Synonyms | | Value | Source |
|---|
| (S)-3-Hydroxyisobutyryl-coenzyme A | HMDB | | 3-Hydroxy-2-methylpropanoyl-CoA | HMDB | | 3-Hydroxy-2-methylpropanoyl-coenzyme A | HMDB | | 3-Hydroxy-2-methylpropionyl-CoA | HMDB | | 3-Hydroxy-2-methylpropionyl-coenzyme A | HMDB | | 3-Hydroxy-isobutyryl-CoA | HMDB | | 3-Hydroxy-isobutyryl-coenzyme A | HMDB | | 3-Hydroxyisobutyryl-CoA | HMDB | | 3-Hydroxyisobutyryl-coenzyme A | HMDB |
|
|---|
| Chemical Formula | C25H42N7O18P3S |
|---|
| Average Molecular Weight | 853.623 |
|---|
| Monoisotopic Molecular Weight | 853.151987801 |
|---|
| IUPAC Name | {[5-(6-amino-9H-purin-9-yl)-4-hydroxy-2-({[hydroxy({[hydroxy(3-hydroxy-3-{[2-({2-[(3-hydroxy-2-methylpropanoyl)sulfanyl]ethyl}carbamoyl)ethyl]carbamoyl}-2,2-dimethylpropoxy)phosphoryl]oxy})phosphoryl]oxy}methyl)oxolan-3-yl]oxy}phosphonic acid |
|---|
| Traditional Name | [5-(6-aminopurin-9-yl)-4-hydroxy-2-({[hydroxy([hydroxy(3-hydroxy-3-{[2-({2-[(3-hydroxy-2-methylpropanoyl)sulfanyl]ethyl}carbamoyl)ethyl]carbamoyl}-2,2-dimethylpropoxy)phosphoryl]oxy)phosphoryl]oxy}methyl)oxolan-3-yl]oxyphosphonic acid |
|---|
| CAS Registry Number | 319440-43-2 |
|---|
| SMILES | CC(CO)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OCC1OC(C(O)C1OP(O)(O)=O)N1C=NC2=C(N)N=CN=C12 |
|---|
| InChI Identifier | InChI=1S/C25H42N7O18P3S/c1-13(8-33)24(38)54-7-6-27-15(34)4-5-28-22(37)19(36)25(2,3)10-47-53(44,45)50-52(42,43)46-9-14-18(49-51(39,40)41)17(35)23(48-14)32-12-31-16-20(26)29-11-30-21(16)32/h11-14,17-19,23,33,35-36H,4-10H2,1-3H3,(H,27,34)(H,28,37)(H,42,43)(H,44,45)(H2,26,29,30)(H2,39,40,41) |
|---|
| InChI Key | WWEOGFZEFHPUAM-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as acyl coas. These are organic compounds containing a coenzyme A substructure linked to an acyl chain. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
| Direct Parent | Acyl CoAs |
|---|
| Alternative Parents | |
|---|
| Substituents | - Coenzyme a or derivatives
- Purine ribonucleoside diphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside 3',5'-bisphosphate
- Ribonucleoside 3'-phosphate
- Pentose-5-phosphate
- Pentose phosphate
- Beta amino acid or derivatives
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Monosaccharide phosphate
- Organic pyrophosphate
- Imidazopyrimidine
- Purine
- Monoalkyl phosphate
- Aminopyrimidine
- Fatty amide
- Monosaccharide
- N-acyl-amine
- N-substituted imidazole
- Organic phosphoric acid derivative
- Imidolactam
- Phosphoric acid ester
- Alkyl phosphate
- Pyrimidine
- Tetrahydrofuran
- Azole
- Heteroaromatic compound
- Imidazole
- Carbothioic s-ester
- Secondary carboxylic acid amide
- Secondary alcohol
- Thiocarboxylic acid ester
- Carboxamide group
- Amino acid or derivatives
- Thiocarboxylic acid or derivatives
- Sulfenyl compound
- Carboxylic acid derivative
- Organoheterocyclic compound
- Oxacycle
- Azacycle
- Hydrocarbon derivative
- Organic nitrogen compound
- Organic oxide
- Organosulfur compound
- Organonitrogen compound
- Primary amine
- Organopnictogen compound
- Carbonyl group
- Organic oxygen compound
- Primary alcohol
- Organooxygen compound
- Alcohol
- Amine
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
| Not Available | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatized |
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| MS | Mass Spectrum (Electron Ionization) | Not Available | 2022-08-06 | Not Available | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 10V, Positive-QTOF | splash10-000i-1912000120-c718f8aabcd5b3328193 | 2015-09-14 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 20V, Positive-QTOF | splash10-000i-1913000000-86eb8b4e6cdee60b559a | 2015-09-14 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 40V, Positive-QTOF | splash10-000i-2911000000-2d6f48b3506835c89ffd | 2015-09-14 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 10V, Negative-QTOF | splash10-00o9-9730140560-056ac34ed9b661751c57 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 20V, Negative-QTOF | splash10-001i-3920110010-37a68eba22e3eecc9757 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 40V, Negative-QTOF | splash10-057i-6900100000-e33bac5e5de19c7b719b | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 10V, Positive-QTOF | splash10-0udi-0000000090-9147721d7b7a39a06164 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 20V, Positive-QTOF | splash10-0fri-1211103790-2282fa97e4f5351f2c0e | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 40V, Positive-QTOF | splash10-0002-0129000000-625efeaf22c52212984a | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 10V, Negative-QTOF | splash10-0udi-0000000090-ff327f166604b01cb3f3 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 20V, Negative-QTOF | splash10-0udr-9800102130-8d5a197de5930d76b37c | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-3-Hydroxyisobutyryl-CoA 40V, Negative-QTOF | splash10-00p1-9103505710-c610ab0387d6ed968440 | 2021-09-22 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum |
|
|---|
| Biological Properties |
|---|
| Cellular Locations | - Extracellular
- Membrane
- Mitochondria
|
|---|
| Biospecimen Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | None |
|---|
| Associated OMIM IDs | None |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB022393 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 86 |
|---|
| KEGG Compound ID | C06000 |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | 47176 |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | 453 |
|---|
| PubChem Compound | 88 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 28259 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | Not Available |
|---|
| MarkerDB ID | Not Available |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Hawes, John W.; Harper, Edwin T. Synthesis of methacrylyl-CoA and (R)- and (S)-3-hydroxyisobutyryl-CoA. Methods in Enzymology (2000), 324(Branched-Chain Amino Acids), 73-79. |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Loupatty FJ, Clayton PT, Ruiter JP, Ofman R, Ijlst L, Brown GK, Thorburn DR, Harris RA, Duran M, Desousa C, Krywawych S, Heales SJ, Wanders RJ: Mutations in the gene encoding 3-hydroxyisobutyryl-CoA hydrolase results in progressive infantile neurodegeneration. Am J Hum Genet. 2007 Jan;80(1):195-9. Epub 2006 Nov 30. [PubMed:17160907 ]
- Lehnert W, Sass JO: Glutaconyl-CoA is the main toxic agent in glutaryl-CoA dehydrogenase deficiency (glutaric aciduria type I). Med Hypotheses. 2005;65(2):330-3. [PubMed:15922108 ]
|
|---|