You are using an unsupported browser. Please upgrade your browser to a newer version to get the best experience on Human Metabolome Database.
| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2006-05-22 15:12:12 UTC |
|---|
| Update Date | 2023-02-21 17:16:28 UTC |
|---|
| HMDB ID | HMDB0002820 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Methylimidazoleacetic acid |
|---|
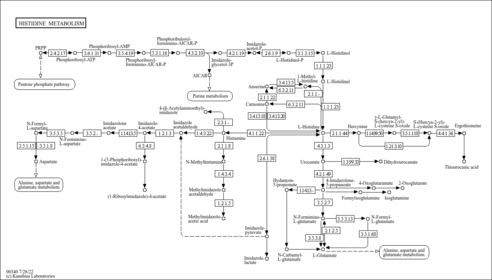
| Description | Methylimidazoleacetic acid is the main metabolite of histamine. This end product of histamine catabolism is formed by N-methylation in the imidazole ring to methylhistamine by histamine methyltransferase (EC 2.1.1.8) and a subsequent oxidative deamination in the side chain by type B monoamine oxidase (EC 1.4.3.4). Based on studies, it is known that as much as 70 to 80 percent of the histamine metabolized in the body is excreted in the urine as methylimidazoleacetic acid. Thus, urinary methylimidazoleacetic acid being the major and specific histamine metabolite is a clear marker of any changes in histamine metabolism in the body. The urinary excretion of methylimidazoleacetic acid is considered a reliable indicator of histamine turnover rate in the body. The excretion of methylimidazoleacetic acid is higher in men than in women. However, this gender difference is abolished when corrected for creatinine excretion. A possible explanation is that basal histamine turnover is related to body size. There is no significant difference in methylimidazoleacetic acid excretion between smokers and non-smokers when analyzing absolute values (mg/24 h). When using methylimidazoleacetic acid values corrected for creatinine excretion female smokers have significantly higher methylimidazoleacetic acid excretion compared to nonsmokers (PMID:11411609 , 7130180 , 10350179 , 10202992 ). |
|---|
| Structure | InChI=1S/C6H8N2O2/c1-8-3-5(7-4-8)2-6(9)10/h3-4H,2H2,1H3,(H,9,10) |
|---|
| Synonyms | | Value | Source |
|---|
| 1,4-Methyl-imidazoleacetic acid | ChEBI | | 1-Methylimidazole-4-acetate | ChEBI | | Methylimidazoleacetate | ChEBI | | tele-Methylimidazoleacetic acid | ChEBI | | 1-Methyl-4-imidazoleacetic acid | Kegg | | 1,4-Methyl-imidazoleacetate | Generator | | 1-Methylimidazole-4-acetic acid | Generator | | tele-Methylimidazoleacetate | Generator | | 1-Methyl-4-imidazoleacetate | Generator | | 1,4-Methylimidazoleacetate | HMDB | | 1-Methyl-1H-imidazole-4-acetate | HMDB | | 1-Methyl-1H-imidazole-4-acetic acid | HMDB | | Methylimidazole acetate | HMDB | | MIAA | HMDB | | N Tau-methylimidazoleacetic acid | HMDB | | Methylimidazoleacetic acid, hydrochloride | MeSH, HMDB |
|
|---|
| Chemical Formula | C6H8N2O2 |
|---|
| Average Molecular Weight | 140.1399 |
|---|
| Monoisotopic Molecular Weight | 140.05857751 |
|---|
| IUPAC Name | 2-(1-methyl-1H-imidazol-4-yl)acetic acid |
|---|
| Traditional Name | methylimidazoleacetic acid |
|---|
| CAS Registry Number | 2625-49-2 |
|---|
| SMILES | CN1C=NC(CC(O)=O)=C1 |
|---|
| InChI Identifier | InChI=1S/C6H8N2O2/c1-8-3-5(7-4-8)2-6(9)10/h3-4H,2H2,1H3,(H,9,10) |
|---|
| InChI Key | ZHCKPJGJQOPTLB-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as imidazolyl carboxylic acids and derivatives. These are organic compounds containing a carboxylic acid chain (of at least 2 carbon atoms) linked to an imidazole ring. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Azoles |
|---|
| Sub Class | Imidazoles |
|---|
| Direct Parent | Imidazolyl carboxylic acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Imidazolyl carboxylic acid derivative
- N-substituted imidazole
- Heteroaromatic compound
- Carboxylic acid derivative
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Azacycle
- Carbonyl group
- Organooxygen compound
- Organonitrogen compound
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organic oxygen compound
- Organic nitrogen compound
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized |
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Methylimidazoleacetic acid GC-MS (1 TMS) | splash10-00kb-3900000000-50b4d0d53d911c99c819 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Methylimidazoleacetic acid GC-MS (Non-derivatized) | splash10-00kb-3900000000-50b4d0d53d911c99c819 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Methylimidazoleacetic acid GC-MS (Non-derivatized) - 70eV, Positive | splash10-0007-9400000000-0a1d81849f89413aebc8 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Methylimidazoleacetic acid GC-MS (1 TMS) - 70eV, Positive | splash10-0072-9200000000-be88458ee3e0fbac72bf | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Methylimidazoleacetic acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-0005-9600000000-bfb5120a71bcb59d0595 | 2012-07-25 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-0002-9000000000-e8be837208f721108cb5 | 2012-07-25 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-00l7-9000000000-30049714cfffac75afa8 | 2012-07-25 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid CE-QTOF-MS system (Agilent 7100 CE + 6550 QTOF) 20V, Positive-QTOF | Not Available | 2019-09-10 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 40V, Positive-QTOF | splash10-0006-9000000000-dedfe29cd079d16b6ab6 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 30V, Positive-QTOF | splash10-0002-9000000000-1eb1e63625c9a88ab50e | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 30V, Positive-QTOF | splash10-0002-9000000000-2091aa9161bf50c2c4a3 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Positive-QTOF | splash10-0002-9100000000-dda19ef742ce86eb3cdc | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 30V, Positive-QTOF | splash10-0002-9000000000-1075d89ff75e92d6eb4d | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 0V, Positive-QTOF | splash10-0006-2900000000-4f104488c46146ea3415 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Positive-QTOF | splash10-0002-9100000000-5d462295a801a6f187f2 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 0V, Positive-QTOF | splash10-0006-2900000000-9dce8cd7f849fb068b41 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Positive-QTOF | splash10-0002-9200000000-c5783540dfb0c4f31a43 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Methylimidazoleacetic acid 20V, Positive-QTOF | splash10-0002-9000000000-b1629f38af1ff6c0c881 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Positive-QTOF | splash10-00di-0900000000-613db7c6c34c85787526 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 20V, Positive-QTOF | splash10-00di-3900000000-b5b0ab7d260b2c465ceb | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 40V, Positive-QTOF | splash10-0zna-9000000000-4d40f26175c89f5ca8f1 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Negative-QTOF | splash10-000i-2900000000-5f3cf75dbdd1a0ff7b4e | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 20V, Negative-QTOF | splash10-0072-5900000000-91359fd775eb604a3d5d | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 40V, Negative-QTOF | splash10-0592-9300000000-24bfa38c15e2d56c8fa6 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Positive-QTOF | splash10-000x-6900000000-ca237928edb79864a607 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 20V, Positive-QTOF | splash10-0002-9300000000-a2a472401cffb11f576f | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 40V, Positive-QTOF | splash10-0uea-9000000000-9a3cd80563596785356a | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 10V, Negative-QTOF | splash10-006t-7900000000-20650405c19d75a6a299 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Methylimidazoleacetic acid 20V, Negative-QTOF | splash10-007a-4900000000-c3539a1cf8d172ae1bb4 | 2021-09-22 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|
| Biological Properties |
|---|
| Cellular Locations | |
|---|
| Biospecimen Locations | - Blood
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Urine
|
|---|
| Tissue Locations | |
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 0.08457 +/- 0.01364 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.08076 +/- 0.01892 uM | Not Specified | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.02 +/- 0.001 (0.021-0.023) uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.022 +/- 0.0021 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.0071 +/- 0.0009 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.50 (0.85-2.3) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.58 (0.48-2.82) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.25 (0.79-3.5) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 2.3 +/- 0.14 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details |
|
|---|
| Abnormal Concentrations |
|---|
| |
| Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Oral cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Breast cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Pancreatic cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Periodontal diseases | | details | | Urine | Detected and Quantified | 10.3 umol/mmol creatinine | Adult (>18 years old) | Both | Mastocytosis | | details |
|
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Perillyl alcohol administration for cancer treatment |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Pancreatic cancer |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Periodontal disease |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Mastocytosis |
|---|
- Martens-Lobenhoffer J, Neumann HJ: Determination of 1-methylhistamine and 1-methylimidazoleacetic acid in human urine as a tool for the diagnosis of mastocytosis. J Chromatogr B Biomed Sci Appl. 1999 Jan 8;721(1):135-40. [PubMed:10027644 ]
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB023069 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 68319 |
|---|
| KEGG Compound ID | C05828 |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | 46587 |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | 3774 |
|---|
| PubChem Compound | 75810 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 1606 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | 3MLDA |
|---|
| MarkerDB ID | MDB00000402 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Download (PDF) |
|---|
| General References | - Martens-Lobenhoffer J, Neumann HJ: Determination of 1-methylhistamine and 1-methylimidazoleacetic acid in human urine as a tool for the diagnosis of mastocytosis. J Chromatogr B Biomed Sci Appl. 1999 Jan 8;721(1):135-40. [PubMed:10027644 ]
- Tsuruta Y, Tomida H, Kohashi K, Ohkura Y: Simultaneous determination of imidazoleacetic acid and N tau- and N pi-methylimidazoleacetic acids in human urine by high-performance liquid chromatography with fluorescence detection. J Chromatogr. 1987 Apr 24;416(1):63-9. [PubMed:3597642 ]
- Khandelwal JK, Hough LB, Pazhenchevsky B, Morrishow AM, Green JP: Presence and measurement of methylimidazoleacetic acids in brain and body fluids. J Biol Chem. 1982 Nov 10;257(21):12815-9. [PubMed:7130180 ]
- Yocum MW, Butterfield JH, Gharib H: Increased plasma calcitonin levels in systemic mast cell disease. Mayo Clin Proc. 1994 Oct;69(10):987-90. [PubMed:7934197 ]
- Prell GD, Khandelwal JK, Burns RS, LeWitt PA, Green JP: Elevated levels of histamine metabolites in cerebrospinal fluid of aging, healthy humans. Compr Gerontol A. 1988 Oct;2(3):114-9. [PubMed:2906817 ]
- Prell GD, Green JP, Kaufmann CA, Khandelwal JK, Morrishow AM, Kirch DG, Linnoila M, Wyatt RJ: Histamine metabolites in cerebrospinal fluid of patients with chronic schizophrenia: their relationships to levels of other aminergic transmitters and ratings of symptoms. Schizophr Res. 1995 Jan;14(2):93-104. [PubMed:7711000 ]
- Granerus G, Roupe G, Swanbeck G: Decreased urinary histamine metabolite after successful PUVA treatment of urticaria pigmentosa. J Invest Dermatol. 1981 Jan;76(1):1-3. [PubMed:7462662 ]
- Swahn CG, Sedvall G: Identification and determination of 1-methylimidazole-4-acetic acid in human cerebrospinal fluid by gas chromatography-mass spectrometry. J Neurochem. 1983 Mar;40(3):688-96. [PubMed:6827268 ]
- Johansson AC, Lonnqvist B, Granerus G: The relationship between body size and the urinary excretion of the main histamine metabolite tele-methylimidazoleacetic acid in man. Inflamm Res. 2001 Apr;50 Suppl 2:S70-1. [PubMed:11411609 ]
- Granerus G, Lonnqvist B, Stenstrom M: No sex difference in the urinary excretion of the histamine metabolite methylimidazoleacetic acid (MeImAA) when corrected for creatinine excretion. Inflamm Res. 1999 Apr;48 Suppl 1:S92-3. [PubMed:10350179 ]
- Granerus G, Lonnqvist B, Wass U: Determination of the histamine metabolite tele-methylimidazoleacetic acid and of creatinine in urine by the same HPLC system. Inflamm Res. 1999 Feb;48(2):75-80. [PubMed:10202992 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
|
|---|