| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-02-21 17:14:33 UTC |
|---|
| HMDB ID | HMDB0000142 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Formic acid |
|---|
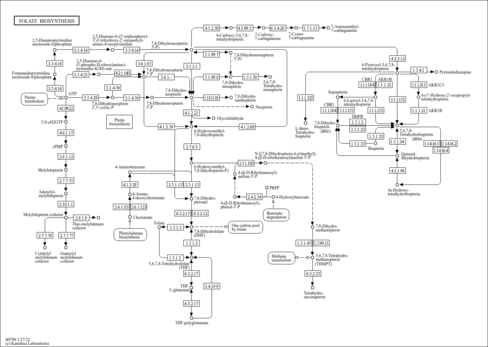
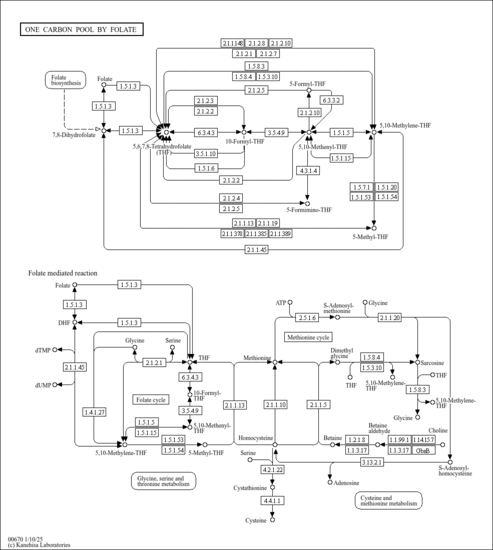
| Description | Formic acid is the simplest carboxylic acid. Formate is an intermediate in normal metabolism. It takes part in the metabolism of one-carbon compounds and its carbon may appear in methyl groups undergoing transmethylation. It is eventually oxidized to carbon dioxide. Formate is typically produced as a byproduct in the production of acetate. It is responsible for both metabolic acidosis and disrupting mitochondrial electron transport and energy production by inhibiting cytochrome oxidase activity, the terminal electron acceptor of the electron transport chain. Cell death from cytochrome oxidase inhibition by formate is believed to result partly from depletion of ATP, reducing energy concentrations so that essential cell functions cannot be maintained. Furthermore, inhibition of cytochrome oxidase by formate may also cause cell death by increased production of cytotoxic reactive oxygen species (ROS) secondary to the blockade of the electron transport chain. In nature, formic acid is found in the stings and bites of many insects of the order Hymenoptera, including bees and ants. The principal use of formic acid is as a preservative and antibacterial agent in livestock feed. When sprayed on fresh hay or other silage, it arrests certain decay processes and causes the feed to retain its nutritive value longer. Urinary formate is produced by Escherichia coli, Pseudomonas aeruginosa, Klebsiella pneumonia, Enterobacter, Acinetobacter, Proteus mirabilis, Citrobacter frundii, Enterococcus faecalis, Streptococcus group B, Staphylococcus saprophyticus (PMID: 22292465 ). |
|---|
| Structure | InChI=1S/CH2O2/c2-1-3/h1H,(H,2,3) |
|---|
| Synonyms | | Value | Source |
|---|
| Acide formique | ChEBI | | Ameisensaeure | ChEBI | | Aminic acid | ChEBI | | Bilorin | ChEBI | | Formylic acid | ChEBI | | H-COOH | ChEBI | | HCO2H | ChEBI | | HCOOH | ChEBI | | Hydrogen carboxylic acid | ChEBI | | Methanoic acid | ChEBI | | Methoic acid | ChEBI | | Aminate | Generator | | Formylate | Generator | | Hydrogen carboxylate | Generator | | Methanoate | Generator | | Methoate | Generator | | Formate | Generator | | Add-F | HMDB | | Ameisensaure | HMDB | | Collo-bueglatt | HMDB | | Collo-didax | HMDB | | Formira | HMDB | | Formisoton | HMDB | | Methanoic acid monomer | HMDB | | Myrmicyl | HMDB | | Sodium formate | HMDB | | Sybest | HMDB | | Wonderbond hardener m 600l | HMDB | | Calcium formate | HMDB | | Cobalt(II) formate dihydrate | HMDB | | Formic acid, aluminum salt | HMDB | | Formic acid, copper salt | HMDB | | Formic acid, cromium (+3) salt | HMDB | | Lithium formate | HMDB | | Ammonium formate | HMDB | | Formic acid, ammonium (4:1) salt | HMDB | | Formic acid, ammonium salt | HMDB | | Formic acid, calcium salt | HMDB | | Formic acid, copper (+2) salt | HMDB | | Formic acid, lead (+2) salt | HMDB | | Formic acid, lead salt | HMDB | | Formic acid, nickel salt | HMDB | | Formic acid, potassium salt | HMDB | | Formic acid, strontium salt | HMDB | | Mafusol | HMDB | | Ammonium tetraformate | HMDB | | Formic acid, 14C-labeled | HMDB | | Formic acid, cobalt (+2) salt | HMDB | | Formic acid, copper, ammonium salt | HMDB | | Formic acid, sodium salt | HMDB | | Formic acid, sodium salt, 14C-labeled | HMDB | | Formic acid, ammonium (2:1) salt | HMDB | | Formic acid, cadmium salt | HMDB | | Formic acid, cesium salt | HMDB | | Formic acid, copper, nickel salt | HMDB | | Formic acid, cromium (+3), sodium (4:1:1) salt | HMDB | | Formic acid, lithium salt | HMDB | | Formic acid, magnesium salt | HMDB | | Formic acid, nickel (+2) salt | HMDB | | Formic acid, rubidium salt | HMDB | | Formic acid, sodium salt, 13C-labeled | HMDB | | Formic acid, thallium (+1) salt | HMDB | | Formic acid, zinc salt | HMDB | | Nickel formate dihydrate | HMDB | | Aluminum formate | HMDB | | Potassium formate | HMDB | | Strontium formate | HMDB | | Lead formate | HMDB | | Nickel formate | HMDB | | Chromic formate | HMDB | | Cobaltous formate | HMDB | | Cupric formate | HMDB | | Magnesium formate | HMDB | | Zinc formate | HMDB |
| Show more...
|---|
| Chemical Formula | CH2O2 |
|---|
| Average Molecular Weight | 46.0254 |
|---|
| Monoisotopic Molecular Weight | 46.005479308 |
|---|
| IUPAC Name | formic acid |
|---|
| Traditional Name | formic acid |
|---|
| CAS Registry Number | 64-18-6 |
|---|
| SMILES | OC=O |
|---|
| InChI Identifier | InChI=1S/CH2O2/c2-1-3/h1H,(H,2,3) |
|---|
| InChI Key | BDAGIHXWWSANSR-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as carboxylic acids. Carboxylic acids are compounds containing a carboxylic acid group with the formula -C(=O)OH. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Carboxylic acids |
|---|
| Direct Parent | Carboxylic acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Monocarboxylic acid or derivatives
- Carboxylic acid
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Not Available | Not Available |
|---|
| Physical Properties |
|---|
| State | Liquid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 8.4 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 1000 mg/mL | Not Available | | LogP | -0.54 | HANSCH,C ET AL. (1995) |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatized| Metabolite | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Formic acid | OC=O | 1434.1 | Standard polar | 33892256 | | Formic acid | OC=O | 488.7 | Standard non polar | 33892256 | | Formic acid | OC=O | 494.5 | Semi standard non polar | 33892256 |
Derivatized | Show more...
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - Formic acid GC-MS (Non-derivatized) - 70eV, Positive | splash10-0002-9000000000-5d27bb312e37a2c8994f | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Formic acid GC-MS (1 TMS) - 70eV, Positive | splash10-0fmi-9200000000-2a89ba98485194acd75a | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Formic acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Formic acid GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-004j-9000000000-2e63b0c1e2e417b0d747 | 2015-03-01 | Not Available | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Formic acid QqQ 4V, positive-QTOF | splash10-0002-9000000000-a8fbddf8ca4197b30013 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Formic acid QqQ 5V, positive-QTOF | splash10-0002-9000000000-98310116f8a2d6a969ce | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Formic acid QqQ 6V, positive-QTOF | splash10-0002-9000000000-ec8753fd9790a3cf0be8 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Formic acid QqQ 7V, positive-QTOF | splash10-0002-9000000000-121d1a025b72a70e412a | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Formic acid QqQ 8V, positive-QTOF | splash10-0002-9000000000-5f1955dee7ab988e86dd | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Formic acid QqQ 9V, positive-QTOF | splash10-0002-9000000000-f8e14272296ee06d19f4 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 10V, Positive-QTOF | splash10-0002-9000000000-092f816e62c8d2f5d56e | 2015-04-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 20V, Positive-QTOF | splash10-0002-9000000000-092f816e62c8d2f5d56e | 2015-04-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 40V, Positive-QTOF | splash10-0002-9000000000-092f816e62c8d2f5d56e | 2015-04-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 10V, Negative-QTOF | splash10-0006-9000000000-eb2207f7400e9144fff7 | 2015-04-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 20V, Negative-QTOF | splash10-0006-9000000000-eb2207f7400e9144fff7 | 2015-04-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 40V, Negative-QTOF | splash10-0006-9000000000-eb2207f7400e9144fff7 | 2015-04-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 10V, Negative-QTOF | splash10-0006-9000000000-d948d5d95ae14e701f57 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 20V, Negative-QTOF | splash10-0006-9000000000-d948d5d95ae14e701f57 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 40V, Negative-QTOF | splash10-0006-9000000000-d948d5d95ae14e701f57 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 10V, Positive-QTOF | splash10-0002-9000000000-92190863fc6ae28a8789 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 20V, Positive-QTOF | splash10-0002-9000000000-92190863fc6ae28a8789 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Formic acid 40V, Positive-QTOF | splash10-0002-9000000000-348f481062f48991a0aa | 2021-09-22 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Mitochondria
- Nucleus
- Endoplasmic reticulum
|
|---|
| Biospecimen Locations | - Blood
- Breast Milk
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Sweat
- Urine
|
|---|
| Tissue Locations | - Brain
- Epidermis
- Fibroblasts
- Kidney
- Liver
- Neuron
- Pancreas
|
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 224.5 +/- 119.8 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 121.7 +/- 97.8 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 106.5 +/- 91.2 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 11.84(29.81) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 23.90-219.5 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 32.8 +/- 13.3 uM | Adult (>18 years old) | Both | Normal | | details | | Breast Milk | Detected and Quantified | 12.2 +/- 14.8 uM | Adult (>18 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 32 +/- 16 uM | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 59 (9-244) uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 38 (3-201) uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 47 (<1-407) uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 35 (7-106) uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 370 +/- 660 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Saliva | Detected and Quantified | 390 +/- 590 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 500 +/- 600 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 600 +/- 750 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 460 +/- 570 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 490 +/- 660 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 340 +/- 520 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 46 (3-244) uM | Adult (>18 years old) | Male | normal | | details | | Saliva | Detected and Quantified | 1-96 uM | Adult (>18 years old) | Male | normal | | details | | Saliva | Detected and Quantified | 96.37 +/- 83.83 uM | Adult (>18 years old) | Both | Normal | | details | | Sweat | Detected and Quantified | < 10 uM | Adult (60 years old) | Male | Normal | | details | | Sweat | Detected and Quantified | < 10 uM | Adult (40 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 44.2 +/- 22.8 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 17 +/- 8.11 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | >0.65 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 43.47-195.63 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.17 (0.07-1.61) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 26.8 (6.9-120.9) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 11.82 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 20.39 +/- 11.84 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 46.198 +/- 30.211 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 27.593 +/- 25.421 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 55.42 (14.60 – 120.70) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details |
| Show more...
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 19.8 +/- 6.8 uM | Adult (>18 years old) | Both | Heart Transplant | | details | | Blood | Detected and Quantified | 171.7 +/- 73.9 uM | Adult (>18 years old) | Both | Methanol poisoning | | details | | Blood | Detected and Quantified | 1630 (543 - 2717) uM | Adult (>18 years old) | Female | Formic acid intoxication | | details | | Blood | Detected and Quantified | 19.1 +/- 21.3 uM | Adult (>18 years old) | Female | Pregnancy with fetuses with trisomy 18 | | details | | Blood | Detected and Quantified | 8.2 +/- 11.2 uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 12.4 (4.5) uM | Adult (>18 years old) | Female | Early preeclampsia | | details | | Blood | Detected and Quantified | 19.3 (14.8) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 600.0 (400.0-17100.0) uM | Adult (>18 years old) | Both | Methanol poisoning | | details | | Blood | Detected and Quantified | 27.0 (13.8) uM | Adult (>18 years old) | Female | Late-onset preeclampsia | | details | | Blood | Detected and Quantified | 29.0 (17.8) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 33.48(33.2) uM | Adult (>18 years old) | Both | Heart failure with preserved ejection fraction | | details | | Blood | Detected and Quantified | 34.2(42.65) uM | Adult (>18 years old) | Both | Heart failure with reduced ejection fraction | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Chronic pancreatitis | | details | | Blood | Detected and Quantified | 3479.9 (628.4) uM | Adult (>18 years old) | Female | Down syndrome pregnancy | | details | | Blood | Detected and Quantified | 3496.6 (620.1) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | ankylosing spondylitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | rheumatoid arthritis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Irritable Bowel Syndrome | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | asymptomatic diverticulosis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | symptomatic uncomplicated diverticular disease | | details | | Urine | Detected and Quantified | 79.1 +/- 27.8 umol/mmol creatinine | Adult (>18 years old) | Both | Methyl formate exposure | | details | | Urine | Detected and Quantified | 0.0006 - 0.0081 umol/mmol creatinine | Adult (>18 years old) | Both | ADPKD | | details | | Urine | Detected and Quantified | 20.0 +/- 11.0 umol/mmol creatinine | Adult (>18 years old) | Both | Lung cancer | | details | | Urine | Detected and Quantified | 31.25 +/- 30.018 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected and Quantified | 64.96 +/- 62.14 umol/mmol creatinine | Adult (>18 years old) | Both | Methanol poisoning | | details | | Urine | Detected and Quantified | 92.45 (20.06 – 524.35) umol/mmol creatinine | Adult (>18 years old) | Both | Type 1 diabetes Mellitus | | details |
| Show more...
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Methanol poisoning |
|---|
- Sejersted OM, Jacobsen D, Ovrebo S, Jansen H: Formate concentrations in plasma from patients poisoned with methanol. Acta Med Scand. 1983;213(2):105-10. [PubMed:6837328 ]
- Baumann K, Angerer J: Occupational chronic exposure to organic solvents. VI. Formic acid concentration in blood and urine as an indicator of methanol exposure. Int Arch Occup Environ Health. 1979 Jan 15;42(3-4):241-9. [PubMed:422265 ]
| | Formic acid intoxication |
|---|
- Moore DF, Bentley AM, Dawling S, Hoare AM, Henry JA: Folinic acid and enhanced renal elimination in formic acid intoxication. J Toxicol Clin Toxicol. 1994;32(2):199-204. [PubMed:8145360 ]
| | Chronic pancreatitis |
|---|
- Zhang L, Jin H, Guo X, Yang Z, Zhao L, Tang S, Mo P, Wu K, Nie Y, Pan Y, Fan D: Distinguishing pancreatic cancer from chronic pancreatitis and healthy individuals by (1)H nuclear magnetic resonance-based metabonomic profiles. Clin Biochem. 2012 Sep;45(13-14):1064-9. doi: 10.1016/j.clinbiochem.2012.05.012. Epub 2012 May 19. [PubMed:22613268 ]
| | Early preeclampsia |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomics and first-trimester prediction of early-onset preeclampsia. J Matern Fetal Neonatal Med. 2012 Oct;25(10):1840-7. doi: 10.3109/14767058.2012.680254. Epub 2012 Apr 28. [PubMed:22494326 ]
| | Pregnancy |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomics and first-trimester prediction of early-onset preeclampsia. J Matern Fetal Neonatal Med. 2012 Oct;25(10):1840-7. doi: 10.3109/14767058.2012.680254. Epub 2012 Apr 28. [PubMed:22494326 ]
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: First-trimester metabolomic detection of late-onset preeclampsia. Am J Obstet Gynecol. 2013 Jan;208(1):58.e1-7. doi: 10.1016/j.ajog.2012.11.003. Epub 2012 Nov 13. [PubMed:23159745 ]
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomic analysis for first-trimester Down syndrome prediction. Am J Obstet Gynecol. 2013 May;208(5):371.e1-8. doi: 10.1016/j.ajog.2012.12.035. Epub 2013 Jan 8. [PubMed:23313728 ]
- Bahado-Singh RO, Akolekar R, Chelliah A, Mandal R, Dong E, Kruger M, Wishart DS, Nicolaides K: Metabolomic analysis for first-trimester trisomy 18 detection. Am J Obstet Gynecol. 2013 Jul;209(1):65.e1-9. doi: 10.1016/j.ajog.2013.03.028. Epub 2013 Mar 25. [PubMed:23535240 ]
| | Late-onset preeclampsia |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: First-trimester metabolomic detection of late-onset preeclampsia. Am J Obstet Gynecol. 2013 Jan;208(1):58.e1-7. doi: 10.1016/j.ajog.2012.11.003. Epub 2012 Nov 13. [PubMed:23159745 ]
| | Irritable bowel syndrome |
|---|
- Hong YS, Hong KS, Park MH, Ahn YT, Lee JH, Huh CS, Lee J, Kim IK, Hwang GS, Kim JS: Metabonomic understanding of probiotic effects in humans with irritable bowel syndrome. J Clin Gastroenterol. 2011 May-Jun;45(5):415-25. doi: 10.1097/MCG.0b013e318207f76c. [PubMed:21494186 ]
| | Diverticular disease |
|---|
- Tursi A, Mastromarino P, Capobianco D, Elisei W, Miccheli A, Capuani G, Tomassini A, Campagna G, Picchio M, Giorgetti G, Fabiocchi F, Brandimarte G: Assessment of Fecal Microbiota and Fecal Metabolome in Symptomatic Uncomplicated Diverticular Disease of the Colon. J Clin Gastroenterol. 2016 Oct;50 Suppl 1:S9-S12. doi: 10.1097/MCG.0000000000000626. [PubMed:27622378 ]
| | Rheumatoid arthritis |
|---|
- Tie-juan ShaoZhi-xing HeZhi-jun XieHai-chang LiMei-jiao WangCheng-ping Wen. Characterization of ankylosing spondylitis and rheumatoid arthritis using 1H NMR-based metabolomics of human fecal extracts. Metabolomics. April 2016, 12:70 [Link]
| | Methyl formate exposure |
|---|
- Berode M, Sethre T, Laubli T, Savolainen H: Urinary methanol and formic acid as indicators of occupational exposure to methyl formate. Int Arch Occup Environ Health. 2000 Aug;73(6):410-4. [PubMed:11007345 ]
| | Lung Cancer |
|---|
- Wishart DS, Knox C, Guo AC, Eisner R, Young N, Gautam B, Hau DD, Psychogios N, Dong E, Bouatra S, Mandal R, Sinelnikov I, Xia J, Jia L, Cruz JA, Lim E, Sobsey CA, Shrivastava S, Huang P, Liu P, Fang L, Peng J, Fradette R, Cheng D, Tzur D, Clements M, Lewis A, De Souza A, Zuniga A, Dawe M, Xiong Y, Clive D, Greiner R, Nazyrova A, Shaykhutdinov R, Li L, Vogel HJ, Forsythe I: HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res. 2009 Jan;37(Database issue):D603-10. doi: 10.1093/nar/gkn810. Epub 2008 Oct 25. [PubMed:18953024 ]
| | Diabetes mellitus type 1 |
|---|
- (). Lorena Ivona ŞTEFAN, Alina NICOLESCU, Simona POPA, Maria MOŢA, Eugenia KOVACS and Calin DELEANU. 1H-NMR URINE METABOLIC PROFILING IN TYPE 1 DIABETES MELLITUS. Rev. Roum. Chim., 2010, 55(11-12), 1033-1037 . .
| | Autosomal dominant polycystic kidney disease |
|---|
- Gronwald W, Klein MS, Zeltner R, Schulze BD, Reinhold SW, Deutschmann M, Immervoll AK, Boger CA, Banas B, Eckardt KU, Oefner PJ: Detection of autosomal dominant polycystic kidney disease by NMR spectroscopic fingerprinting of urine. Kidney Int. 2011 Jun;79(11):1244-53. doi: 10.1038/ki.2011.30. Epub 2011 Mar 9. [PubMed:21389975 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| Associated OMIM IDs | - 1678 (Chronic pancreatitis)
- 180300 (Rheumatoid arthritis)
- 211980 (Lung Cancer)
- 222100 (Diabetes mellitus type 1)
- 601313 (Autosomal dominant polycystic kidney disease)
- 610247 (Eosinophilic esophagitis)
|
|---|
| External Links |
|---|
| DrugBank ID | DB01942 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB012804 |
|---|
| KNApSAcK ID | C00001182 |
|---|
| Chemspider ID | Not Available |
|---|
| KEGG Compound ID | C00058 |
|---|
| BioCyc ID | FORMATE |
|---|
| BiGG ID | Not Available |
|---|
| Wikipedia Link | Formic_acid |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 284 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 30751 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | FOR |
|---|
| MarkerDB ID | MDB00000068 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Finholt, Albert E.; Jacobson, Eugene C. The reduction of carbon dioxide to formic acid with lithium aluminum hydride. Journal of the American Chemical Society (1952), 74 3943-4. |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Nicholson JK, Foxall PJ, Spraul M, Farrant RD, Lindon JC: 750 MHz 1H and 1H-13C NMR spectroscopy of human blood plasma. Anal Chem. 1995 Mar 1;67(5):793-811. [PubMed:7762816 ]
- Dunne VG, Bhattachayya S, Besser M, Rae C, Griffin JL: Metabolites from cerebrospinal fluid in aneurysmal subarachnoid haemorrhage correlate with vasospasm and clinical outcome: a pattern-recognition 1H NMR study. NMR Biomed. 2005 Feb;18(1):24-33. [PubMed:15455468 ]
- Bales JR, Higham DP, Howe I, Nicholson JK, Sadler PJ: Use of high-resolution proton nuclear magnetic resonance spectroscopy for rapid multi-component analysis of urine. Clin Chem. 1984 Mar;30(3):426-32. [PubMed:6321058 ]
- Ohmori S, Sumii I, Toyonaga Y, Nakata K, Kawase M: High-performance liquid chromatographic determination of formate as benzimidazole in biological samples. J Chromatogr. 1988 Apr 8;426(1):15-24. [PubMed:3384868 ]
- Dal Pra I, Chiarini A, Boschi A, Freddi G, Armato U: Novel dermo-epidermal equivalents on silk fibroin-based formic acid-crosslinked three-dimensional nonwoven devices with prospective applications in human tissue engineering/regeneration/repair. Int J Mol Med. 2006 Aug;18(2):241-7. [PubMed:16820930 ]
- Igeta Y, Kawarabayashi T, Sato M, Yamada N, Matsubara E, Ishiguro K, Kanai M, Tomidokoro Y, Osuga J, Okamoto K, Hirai S, Shoji M: Apolipoprotein E accumulates with the progression of A beta deposition in transgenic mice. J Neuropathol Exp Neurol. 1997 Nov;56(11):1228-35. [PubMed:9370233 ]
- Kerns W 2nd, Tomaszewski C, McMartin K, Ford M, Brent J: Formate kinetics in methanol poisoning. J Toxicol Clin Toxicol. 2002;40(2):137-43. [PubMed:12126185 ]
- Nagasawa H, Wada M, Koyama S, Kawanami T, Kurita K, Kato T: [A case of methanol intoxication with optic neuropathy visualized on STIR sequence of MR images]. Rinsho Shinkeigaku. 2005 Jul;45(7):527-30. [PubMed:16119839 ]
- Foulon V, Sniekers M, Huysmans E, Asselberghs S, Mahieu V, Mannaerts GP, Van Veldhoven PP, Casteels M: Breakdown of 2-hydroxylated straight chain fatty acids via peroxisomal 2-hydroxyphytanoyl-CoA lyase: a revised pathway for the alpha-oxidation of straight chain fatty acids. J Biol Chem. 2005 Mar 18;280(11):9802-12. Epub 2005 Jan 11. [PubMed:15644336 ]
- Iwamoto N, Nishiyama E, Ohwada J, Arai H: Distribution of amyloid deposits in the cerebral white matter of the Alzheimer's disease brain: relationship to blood vessels. Acta Neuropathol. 1997 Apr;93(4):334-40. [PubMed:9113198 ]
- Ferrari LA, Arado MG, Nardo CA, Giannuzzi L: Post-mortem analysis of formic acid disposition in acute methanol intoxication. Forensic Sci Int. 2003 Apr 23;133(1-2):152-8. [PubMed:12742704 ]
- Tasaka Y, Nakaya F, Matsumoto H, Iwamoto Y, Omori Y: Pancreatic amylin content in human diabetic subjects and its relation to diabetes. Pancreas. 1995 Oct;11(3):303-8. [PubMed:8577686 ]
- D'Andrea MR, Reiser PA, Polkovitch DA, Gumula NA, Branchide B, Hertzog BM, Schmidheiser D, Belkowski S, Gastard MC, Andrade-Gordon P: The use of formic acid to embellish amyloid plaque detection in Alzheimer's disease tissues misguides key observations. Neurosci Lett. 2003 May 15;342(1-2):114-8. [PubMed:12727331 ]
- Berode M, Sethre T, Laubli T, Savolainen H: Urinary methanol and formic acid as indicators of occupational exposure to methyl formate. Int Arch Occup Environ Health. 2000 Aug;73(6):410-4. [PubMed:11007345 ]
- Lehmann P, Kligman AM: In vivo removal of the horny layer with formic acid. Br J Dermatol. 1983 Sep;109(3):313-20. [PubMed:6615718 ]
- Bloomer JC, Clarke SE, Chenery RJ: Determination of P4501A2 activity in human liver microsomes using [3-14C-methyl]caffeine. Xenobiotica. 1995 Sep;25(9):917-27. [PubMed:8553685 ]
- Grady S, Osterloh J: Improved enzymic assay for serum formate with colorimetric endpoint. J Anal Toxicol. 1986 Jan-Feb;10(1):1-5. [PubMed:3754027 ]
- Baumann K, Angerer J: Occupational chronic exposure to organic solvents. VI. Formic acid concentration in blood and urine as an indicator of methanol exposure. Int Arch Occup Environ Health. 1979 Jan 15;42(3-4):241-9. [PubMed:422265 ]
- Ferry DG, Temple WA, McQueen EG: Methanol monitoring. Comparison of urinary methanol concentration with formic acid excretion rate as a measure of occupational exposure. Int Arch Occup Environ Health. 1980;47(2):155-63. [PubMed:7440001 ]
- Gupta A, Dwivedi M, Mahdi AA, Khetrapal CL, Bhandari M: Broad identification of bacterial type in urinary tract infection using (1)h NMR spectroscopy. J Proteome Res. 2012 Mar 2;11(3):1844-54. doi: 10.1021/pr2010692. Epub 2012 Jan 31. [PubMed:22292465 ]
| Show more...
|---|