| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-05-30 20:55:52 UTC |
|---|
| HMDB ID | HMDB0000695 |
|---|
| Secondary Accession Numbers | - HMDB0000762
- HMDB00695
- HMDB00762
|
|---|
| Metabolite Identification |
|---|
| Common Name | Ketoleucine |
|---|
| Description | Ketoleucine is an abnormal metabolite that arises from the incomplete breakdown of branched-chain amino acids. Ketoleucine is both a neurotoxin and a metabotoxin. A neurotoxin causes damage to nerve cells and nerve tissues. A metabotoxin is an endogenously produced metabolite that causes adverse health effects at chronically high levels. Chronically high levels of ketoleucine are associated with maple syrup urine disease (MSUD). MSUD is a metabolic disorder caused by a deficiency of the branched-chain alpha-keto acid dehydrogenase complex (BCKDC), leading to a buildup of the branched-chain amino acids (leucine, isoleucine, and valine) and their toxic by-products (ketoacids) in the blood and urine. The symptoms of MSUD often show in infancy and lead to severe brain damage if untreated. MSUD may also present later depending on the severity of the disease. If left untreated in older individuals, during times of metabolic crisis, symptoms of the condition include uncharacteristically inappropriate, extreme, or erratic behaviour and moods, hallucinations, anorexia, weight loss, anemia, diarrhea, vomiting, dehydration, lethargy, oscillating hypertonia and hypotonia, ataxia, seizures, hypoglycemia, ketoacidosis, opisthotonus, pancreatitis, rapid neurological decline, and coma. In maple syrup urine disease, the brain concentration of branched-chain ketoacids can increase 10- to 20-fold. This leads to a depletion of glutamate and a consequent reduction in the concentration of brain glutamine, aspartate, alanine, and other amino acids. The result is a compromise of energy metabolism because of a failure of the malate-aspartate shuttle and a diminished rate of protein synthesis (PMID: 15930465 ). |
|---|
| Structure | InChI=1S/C6H10O3/c1-4(2)3-5(7)6(8)9/h4H,3H2,1-2H3,(H,8,9) |
|---|
| Synonyms | | Value | Source |
|---|
| 2-oxo-4-METHYLPENTANOIC ACID | ChEBI | | 2-Oxoisocaproate | ChEBI | | 4-Methyl-2-oxopentanoate | ChEBI | | 4-Methyl-2-oxovaleric acid | ChEBI | | alpha-Ketoisocaproic acid | ChEBI | | 2-oxo-4-METHYLPENTANOate | Generator | | 2-Oxoisocaproic acid | Generator | | 4-Methyl-2-oxopentanoic acid | Generator | | 4-Methyl-2-oxovalerate | Generator | | a-Ketoisocaproate | Generator | | a-Ketoisocaproic acid | Generator | | alpha-Ketoisocaproate | Generator | | Α-ketoisocaproate | Generator | | Α-ketoisocaproic acid | Generator | | 2-Keto-4-methylvalerate | HMDB | | 2-Keto-4-methylvaleric acid | HMDB | | 2-Ketoisocaproate | HMDB | | 2-Ketoisocaproic acid | HMDB | | 2-oxo-4-Methylvalerate | HMDB | | 2-oxo-4-Methylvaleric acid | HMDB | | 2-Oxoleucine | HMDB | | 4-Methyl-2-oxo-valerate | HMDB | | 4-Methyl-2-oxo-valeric acid | HMDB | | a-Ketoisocapronate | HMDB | | a-Ketoisocapronic acid | HMDB | | a-Oxoisocaproate | HMDB | | a-Oxoisocaproic acid | HMDB | | alpha-Keto-isocaproate | HMDB | | alpha-Keto-isocaproic acid | HMDB | | alpha-Ketoisocapronate | HMDB | | alpha-Ketoisocapronic acid | HMDB | | alpha-Oxoisocaproate | HMDB | | alpha-Oxoisocaproic acid | HMDB | | Ketoisocaproate | HMDB | | Ketoisocaproic acid | HMDB | | Methyloxovalerate | HMDB | | Methyloxovaleric acid | HMDB | | Oxoisocaproate | HMDB | | Oxoisocaproic acid | HMDB | | Keto-leucine | HMDB |
|
|---|
| Chemical Formula | C6H10O3 |
|---|
| Average Molecular Weight | 130.1418 |
|---|
| Monoisotopic Molecular Weight | 130.062994186 |
|---|
| IUPAC Name | 4-methyl-2-oxopentanoic acid |
|---|
| Traditional Name | ketoisocaproate |
|---|
| CAS Registry Number | 816-66-0 |
|---|
| SMILES | CC(C)CC(=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H10O3/c1-4(2)3-5(7)6(8)9/h4H,3H2,1-2H3,(H,8,9) |
|---|
| InChI Key | BKAJNAXTPSGJCU-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as short-chain keto acids and derivatives. These are keto acids with an alkyl chain the contains less than 6 carbon atoms. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Keto acids and derivatives |
|---|
| Sub Class | Short-chain keto acids and derivatives |
|---|
| Direct Parent | Short-chain keto acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Branched fatty acid
- Methyl-branched fatty acid
- Short-chain keto acid
- Alpha-keto acid
- Fatty acyl
- Alpha-hydroxy ketone
- Ketone
- Carboxylic acid derivative
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Carbonyl group
- Hydrocarbon derivative
- Organic oxygen compound
- Organooxygen compound
- Organic oxide
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Liquid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 8 - 10 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 32 mg/mL | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections| Adduct Type | Data Source | CCS Value (Å2) | Reference |
|---|
| [M-H]- | Astarita_neg | 125.3 | 30932474 |
|
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Ketoleucine,1TMS,isomer #1 | CC(C)CC(=O)C(=O)O[Si](C)(C)C | 1148.8 | Semi standard non polar | 33892256 | | Ketoleucine,1TMS,isomer #2 | CC(C)C=C(O[Si](C)(C)C)C(=O)O | 1235.4 | Semi standard non polar | 33892256 | | Ketoleucine,2TMS,isomer #1 | CC(C)C=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1276.7 | Semi standard non polar | 33892256 | | Ketoleucine,2TMS,isomer #1 | CC(C)C=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1255.8 | Standard non polar | 33892256 | | Ketoleucine,2TMS,isomer #1 | CC(C)C=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1283.4 | Standard polar | 33892256 | | Ketoleucine,1TBDMS,isomer #1 | CC(C)CC(=O)C(=O)O[Si](C)(C)C(C)(C)C | 1369.9 | Semi standard non polar | 33892256 | | Ketoleucine,1TBDMS,isomer #2 | CC(C)C=C(O[Si](C)(C)C(C)(C)C)C(=O)O | 1468.8 | Semi standard non polar | 33892256 | | Ketoleucine,2TBDMS,isomer #1 | CC(C)C=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1709.4 | Semi standard non polar | 33892256 | | Ketoleucine,2TBDMS,isomer #1 | CC(C)C=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1656.5 | Standard non polar | 33892256 | | Ketoleucine,2TBDMS,isomer #1 | CC(C)C=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1617.1 | Standard polar | 33892256 |
|
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-000i-9400000000-49d055f04bfe1877ac5b | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-029j-8930000000-c6c54bf6ee2995a2195b | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0002-1910000000-0ba38257e25873312809 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-MS (2 TMS) | splash10-0a5c-4930000000-61247369a805fa2a77a7 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-MS (1 MEOX; 1 TMS) | splash10-000i-9510000000-8bfc5a54e29f65edf361 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-MS (1 MEOX; 1 TMS) | splash10-00lj-9620000000-aa8409ceb2359d903f38 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Non-derivatized) | splash10-000i-9400000000-49d055f04bfe1877ac5b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Non-derivatized) | splash10-029j-8930000000-c6c54bf6ee2995a2195b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Non-derivatized) | splash10-0002-1910000000-0ba38257e25873312809 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-MS (Non-derivatized) | splash10-0a5c-4930000000-61247369a805fa2a77a7 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-MS (Non-derivatized) | splash10-000i-9510000000-8bfc5a54e29f65edf361 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-MS (Non-derivatized) | splash10-00lj-9620000000-aa8409ceb2359d903f38 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Non-derivatized) | splash10-00ks-7930000000-4e9cfd715fa884d30e7f | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Ketoleucine GC-EI-TOF (Non-derivatized) | splash10-000i-9500000000-709ebcb57f006703c269 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (Non-derivatized) - 70eV, Positive | splash10-0006-9000000000-e410bce23706f826f8dd | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (1 TMS) - 70eV, Positive | splash10-0006-9400000000-c4f06cea1e27a2b24660 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Ketoleucine GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-0006-9000000000-bed7ac074a60210971e3 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-014i-9200000000-2fa08d2b4df406cf63ea | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-014j-9000000000-045be8e8dca3f93d45ba | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-004i-0900000000-237fa8ad6bfb929be31e | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-004r-9700000000-8d38d24fce78b1ff13e6 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-0a59-9000000000-759cfc0817917283d1a5 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-0a4l-9000000000-2a068ada71fbe0a5a988 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative-QTOF | splash10-0a4l-9000000000-58e61e31f3c96d84d799 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ , negative-QTOF | splash10-004i-0900000000-237fa8ad6bfb929be31e | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ , negative-QTOF | splash10-004r-9700000000-8d38d24fce78b1ff13e6 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ , negative-QTOF | splash10-0a59-9000000000-759cfc0817917283d1a5 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ , negative-QTOF | splash10-0a4l-9000000000-2a068ada71fbe0a5a988 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-QQ , negative-QTOF | splash10-0a4l-9000000000-58e61e31f3c96d84d799 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-ITFT , negative-QTOF | splash10-004i-0900000000-556a0d1cf568609752d1 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine LC-ESI-IT , negative-QTOF | splash10-004r-5900000000-20f6923c8ed45e9fa088 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine 35V, Negative-QTOF | splash10-004i-0900000000-b66d521478c92e2d77d0 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine 10V, Negative-QTOF | splash10-004i-0900000000-9c3a698d9f28af574ef6 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine 40V, Negative-QTOF | splash10-03di-9000000000-47ced70de4860e900283 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Ketoleucine 20V, Negative-QTOF | splash10-03xr-7900000000-0461a5a56e36a03fe522 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Ketoleucine 10V, Positive-QTOF | splash10-08gr-7900000000-31021fda89eb6535185f | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Ketoleucine 20V, Positive-QTOF | splash10-0btm-9200000000-3bf5df4b557ef87267f4 | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Ketoleucine 40V, Positive-QTOF | splash10-0a4i-9000000000-37839bf70647d24b94a4 | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Ketoleucine 10V, Negative-QTOF | splash10-004i-4900000000-796467909d1d4c087a07 | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Ketoleucine 20V, Negative-QTOF | splash10-01ri-9400000000-0abcd021721f32c10e42 | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Ketoleucine 40V, Negative-QTOF | splash10-015l-9000000000-5c33a47dfeb09a182983 | 2015-05-27 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane
- Mitochondria
|
|---|
| Biospecimen Locations | - Blood
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Urine
|
|---|
| Tissue Locations | - Neuron
- Placenta
- Prostate
- Skeletal Muscle
|
|---|
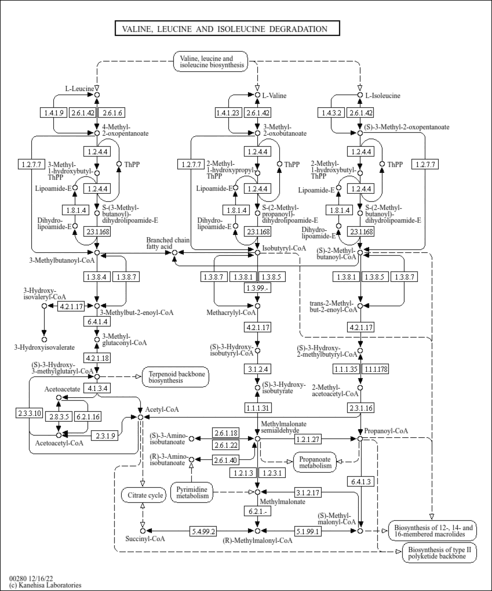
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 28.0 (0.0-58.0) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 33.5 +/- 8.2 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 25-28 uM | Adolescent (13-18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 18 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 5.0 +/- 4.4 uM | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 0.944 +/- 0.442 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 1.24 +/- 0.728 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 0-2 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.48-0.86 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 1.623 +/- 1.21 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.48-0.72 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 0-0.1 umol/mmol creatinine | Infant (0-1 year old) | Female | Normal | | details | | Urine | Detected and Quantified | 0-0.2 umol/mmol creatinine | Infant (0-1 year old) | Male | Normal | | details | | Urine | Detected and Quantified | 0-0.7 umol/mmol creatinine | Infant (0-1 year old) | Female | Normal | | details | | Urine | Detected and Quantified | 0.58 (0.39-0.87) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.13 (0.02-0.45) umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 0.13 (0.03-0.34) umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details |
|
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 1918 uM | Children (1-13 years old) | Female | Maple syrup urine disease | | details | | Blood | Detected and Quantified | 0 uM | Adult (>18 years old) | Female | Branched-chain Keto Acid Dehydrogenase Kinase Deficiency | | details | | Blood | Detected and Quantified | 0 uM | Adolescent (13-18 years old) | Female | Branched-chain Keto Acid Dehydrogenase Kinase Deficiency | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 483 uM | Children (1-13 years old) | Female | Maple syrup urine disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Gout | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | ankylosing spondylitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | rheumatoid arthritis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Attachment loss | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Missing teeth | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Periodontal Probing Depth | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Tooth Decay | | details | | Urine | Detected and Quantified | 1.407 +/- 0.928 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details |
|
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Maple syrup urine disease |
|---|
- Wendel, U., Becker, K., Przyrembel, H. et al. (1980). Peritoneal dialysis in maple-syrup-urine disease: Studies on branched-chain amino and keto acids. Eur J Pediatr (1980) 134: 57. https://doi.org/10.1007/BF00442404. Eur J Pediatr.
| | Branched-chain Keto Acid Dehydrogenase Kinase Deficiency |
|---|
- Novarino G, El-Fishawy P, Kayserili H, Meguid NA, Scott EM, Schroth J, Silhavy JL, Kara M, Khalil RO, Ben-Omran T, Ercan-Sencicek AG, Hashish AF, Sanders SJ, Gupta AR, Hashem HS, Matern D, Gabriel S, Sweetman L, Rahimi Y, Harris RA, State MW, Gleeson JG: Mutations in BCKD-kinase lead to a potentially treatable form of autism with epilepsy. Science. 2012 Oct 19;338(6105):394-7. doi: 10.1126/science.1224631. Epub 2012 Sep 6. [PubMed:22956686 ]
| | Colorectal cancer |
|---|
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
| | Gout |
|---|
- Shao T, Shao L, Li H, Xie Z, He Z, Wen C: Combined Signature of the Fecal Microbiome and Metabolome in Patients with Gout. Front Microbiol. 2017 Feb 21;8:268. doi: 10.3389/fmicb.2017.00268. eCollection 2017. [PubMed:28270806 ]
| | Rheumatoid arthritis |
|---|
- Tie-juan ShaoZhi-xing HeZhi-jun XieHai-chang LiMei-jiao WangCheng-ping Wen. Characterization of ankylosing spondylitis and rheumatoid arthritis using 1H NMR-based metabolomics of human fecal extracts. Metabolomics. April 2016, 12:70 [Link]
| | Attachment loss |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Missing teeth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Periodontal Probing Depth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Tooth Decay |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| Associated OMIM IDs | - 248600 (Maple syrup urine disease)
- 614923 (Branched-chain Keto Acid Dehydrogenase Kinase Deficiency)
- 114500 (Colorectal cancer)
- 138900 (Gout)
- 180300 (Rheumatoid arthritis)
- 610247 (Eosinophilic esophagitis)
|
|---|
| External Links |
|---|
| DrugBank ID | DB03229 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB030510 |
|---|
| KNApSAcK ID | C00019677 |
|---|
| Chemspider ID | 69 |
|---|
| KEGG Compound ID | C00233 |
|---|
| BioCyc ID | 2K-4CH3-PENTANOATE |
|---|
| BiGG ID | 34334 |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | 5663 |
|---|
| PubChem Compound | 70 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 48430 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | 4MOP |
|---|
| MarkerDB ID | MDB00000223 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Tabuchi, Takeshi; Abe, Matazo. Fermentative preparation of a-ketoisocaproic acid. Jpn. Tokkyo Koho (1970), 3 pp. |
|---|
| Material Safety Data Sheet (MSDS) | Download (PDF) |
|---|
| General References | - Martin PM, Gopal E, Ananth S, Zhuang L, Itagaki S, Prasad BM, Smith SB, Prasad PD, Ganapathy V: Identity of SMCT1 (SLC5A8) as a neuron-specific Na+-coupled transporter for active uptake of L-lactate and ketone bodies in the brain. J Neurochem. 2006 Jul;98(1):279-88. [PubMed:16805814 ]
- Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Yudkoff M, Daikhin Y, Nissim I, Horyn O, Luhovyy B, Lazarow A, Nissim I: Brain amino acid requirements and toxicity: the example of leucine. J Nutr. 2005 Jun;135(6 Suppl):1531S-8S. [PubMed:15930465 ]
- Chow LS, Albright RC, Bigelow ML, Toffolo G, Cobelli C, Nair KS: Mechanism of insulin's anabolic effect on muscle: measurements of muscle protein synthesis and breakdown using aminoacyl-tRNA and other surrogate measures. Am J Physiol Endocrinol Metab. 2006 Oct;291(4):E729-36. Epub 2006 May 16. [PubMed:16705065 ]
- Wang Y, Holmes E, Nicholson JK, Cloarec O, Chollet J, Tanner M, Singer BH, Utzinger J: Metabonomic investigations in mice infected with Schistosoma mansoni: an approach for biomarker identification. Proc Natl Acad Sci U S A. 2004 Aug 24;101(34):12676-81. Epub 2004 Aug 16. [PubMed:15314235 ]
- Mitch WE, Walser M, Sapir DG: Nitrogen sparing induced by leucine compared with that induced by its keto analogue, alpha-ketoisocaproate, in fasting obese man. J Clin Invest. 1981 Feb;67(2):553-62. [PubMed:7462428 ]
- Sgaravatti AM, Rosa RB, Schuck PF, Ribeiro CA, Wannmacher CM, Wyse AT, Dutra-Filho CS, Wajner M: Inhibition of brain energy metabolism by the alpha-keto acids accumulating in maple syrup urine disease. Biochim Biophys Acta. 2003 Nov 20;1639(3):232-8. [PubMed:14636955 ]
- Schadewaldt P, Hammen HW, Ott AC, Wendel U: Renal clearance of branched-chain L-amino and 2-oxo acids in maple syrup urine disease. J Inherit Metab Dis. 1999 Aug;22(6):706-22. [PubMed:10472531 ]
- Hachey DL, Patterson BW, Reeds PJ, Elsas LJ: Isotopic determination of organic keto acid pentafluorobenzyl esters in biological fluids by negative chemical ionization gas chromatography/mass spectrometry. Anal Chem. 1991 May 1;63(9):919-23. [PubMed:1858984 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
|
|---|