| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-02-21 17:15:21 UTC |
|---|
| HMDB ID | HMDB0000965 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Hypotaurine |
|---|
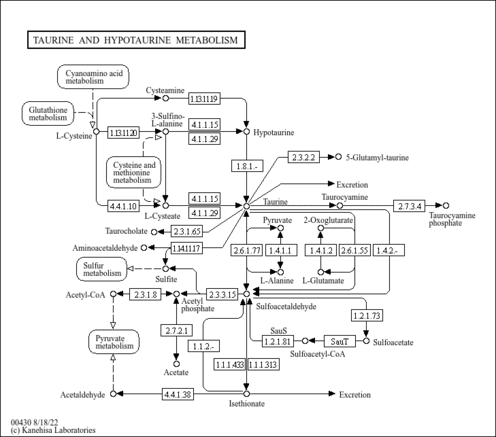
| Description | Hypotaurine belongs to the class of organic compounds known as sulfinic acids. Sulfinic acids are compounds containing a sulfinic acid functional group, with the general structure RS(=O)OH (R = organyl, not H). Hypotaurine exists in all living species, ranging from bacteria to humans. Within humans, hypotaurine participates in a number of enzymatic reactions. In particular, hypotaurine can be biosynthesized from cysteamine; which is catalyzed by the enzyme 2-aminoethanethiol dioxygenase. In addition, hypotaurine can be biosynthesized from 3-sulfinoalanine through its interaction with the enzyme cysteine sulfinic acid decarboxylase. In humans, hypotaurine is involved in taurine and hypotaurine metabolism. |
|---|
| Structure | InChI=1S/C2H7NO2S/c3-1-2-6(4)5/h1-3H2,(H,4,5) |
|---|
| Synonyms | | Value | Source |
|---|
| 2-Aminoethanesulfinic acid | ChEBI | | 2-Aminoethanesulfinate | Generator | | 2-Aminoethanesulphinate | Generator | | 2-Aminoethanesulphinic acid | Generator | | 2-Amino-ethanesulfinate | HMDB | | 2-Amino-ethanesulfinic acid | HMDB | | 2-Aminoethylsulfinate | HMDB | | 2-Aminoethylsulfinic acid | HMDB | | Cystaminesulfinate | HMDB | | Cystaminesulfinic acid | HMDB |
|
|---|
| Chemical Formula | C2H7NO2S |
|---|
| Average Molecular Weight | 109.147 |
|---|
| Monoisotopic Molecular Weight | 109.019749163 |
|---|
| IUPAC Name | 2-aminoethane-1-sulfinic acid |
|---|
| Traditional Name | hypotaurine |
|---|
| CAS Registry Number | 300-84-5 |
|---|
| SMILES | NCCS(O)=O |
|---|
| InChI Identifier | InChI=1S/C2H7NO2S/c3-1-2-6(4)5/h1-3H2,(H,4,5) |
|---|
| InChI Key | VVIUBCNYACGLLV-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as sulfinic acids. Sulfinic acids are compounds containing a sulfinic acid functional group, with the general structure RS(=O)OH (R = organyl, not H). |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Sulfinic acids and derivatives |
|---|
| Sub Class | Sulfinic acids |
|---|
| Direct Parent | Sulfinic acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Sulfinic acid
- Alkanesulfinic acid
- Alkanesulfinic acid or derivatives
- Organic nitrogen compound
- Organic oxygen compound
- Organopnictogen compound
- Organic oxide
- Hydrocarbon derivative
- Primary amine
- Organosulfur compound
- Organonitrogen compound
- Primary aliphatic amine
- Amine
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Hypotaurine,1TMS,isomer #1 | C[Si](C)(C)OS(=O)CCN | 1209.8 | Semi standard non polar | 33892256 | | Hypotaurine,1TMS,isomer #1 | C[Si](C)(C)OS(=O)CCN | 1330.2 | Standard non polar | 33892256 | | Hypotaurine,1TMS,isomer #1 | C[Si](C)(C)OS(=O)CCN | 1878.9 | Standard polar | 33892256 | | Hypotaurine,1TMS,isomer #2 | C[Si](C)(C)NCCS(=O)O | 1322.5 | Semi standard non polar | 33892256 | | Hypotaurine,1TMS,isomer #2 | C[Si](C)(C)NCCS(=O)O | 1393.6 | Standard non polar | 33892256 | | Hypotaurine,1TMS,isomer #2 | C[Si](C)(C)NCCS(=O)O | 1977.0 | Standard polar | 33892256 | | Hypotaurine,2TMS,isomer #1 | C[Si](C)(C)NCCS(=O)O[Si](C)(C)C | 1359.8 | Semi standard non polar | 33892256 | | Hypotaurine,2TMS,isomer #1 | C[Si](C)(C)NCCS(=O)O[Si](C)(C)C | 1590.1 | Standard non polar | 33892256 | | Hypotaurine,2TMS,isomer #1 | C[Si](C)(C)NCCS(=O)O[Si](C)(C)C | 1495.3 | Standard polar | 33892256 | | Hypotaurine,2TMS,isomer #2 | C[Si](C)(C)N(CCS(=O)O)[Si](C)(C)C | 1515.5 | Semi standard non polar | 33892256 | | Hypotaurine,2TMS,isomer #2 | C[Si](C)(C)N(CCS(=O)O)[Si](C)(C)C | 1688.9 | Standard non polar | 33892256 | | Hypotaurine,2TMS,isomer #2 | C[Si](C)(C)N(CCS(=O)O)[Si](C)(C)C | 1812.3 | Standard polar | 33892256 | | Hypotaurine,3TMS,isomer #1 | C[Si](C)(C)OS(=O)CCN([Si](C)(C)C)[Si](C)(C)C | 1558.5 | Semi standard non polar | 33892256 | | Hypotaurine,3TMS,isomer #1 | C[Si](C)(C)OS(=O)CCN([Si](C)(C)C)[Si](C)(C)C | 1809.8 | Standard non polar | 33892256 | | Hypotaurine,3TMS,isomer #1 | C[Si](C)(C)OS(=O)CCN([Si](C)(C)C)[Si](C)(C)C | 1513.7 | Standard polar | 33892256 | | Hypotaurine,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OS(=O)CCN | 1425.6 | Semi standard non polar | 33892256 | | Hypotaurine,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OS(=O)CCN | 1628.1 | Standard non polar | 33892256 | | Hypotaurine,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OS(=O)CCN | 1958.6 | Standard polar | 33892256 | | Hypotaurine,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)NCCS(=O)O | 1563.3 | Semi standard non polar | 33892256 | | Hypotaurine,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)NCCS(=O)O | 1698.3 | Standard non polar | 33892256 | | Hypotaurine,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)NCCS(=O)O | 2030.7 | Standard polar | 33892256 | | Hypotaurine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)NCCS(=O)O[Si](C)(C)C(C)(C)C | 1791.1 | Semi standard non polar | 33892256 | | Hypotaurine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)NCCS(=O)O[Si](C)(C)C(C)(C)C | 2135.9 | Standard non polar | 33892256 | | Hypotaurine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)NCCS(=O)O[Si](C)(C)C(C)(C)C | 1738.4 | Standard polar | 33892256 | | Hypotaurine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N(CCS(=O)O)[Si](C)(C)C(C)(C)C | 1945.0 | Semi standard non polar | 33892256 | | Hypotaurine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N(CCS(=O)O)[Si](C)(C)C(C)(C)C | 2172.9 | Standard non polar | 33892256 | | Hypotaurine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N(CCS(=O)O)[Si](C)(C)C(C)(C)C | 1915.8 | Standard polar | 33892256 | | Hypotaurine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OS(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2202.8 | Semi standard non polar | 33892256 | | Hypotaurine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OS(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2524.2 | Standard non polar | 33892256 | | Hypotaurine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OS(=O)CCN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 1873.8 | Standard polar | 33892256 |
|
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0udr-0900000000-6d1ec3a7649c0e7a704b | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0f79-0900000000-468e3da761e4d86223b1 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0fki-5900000000-0569fb6d8502d22d2b7e | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-MS (3 TMS) | splash10-0f79-0900000000-457c0f4ff1563d09a208 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine EI-B (Non-derivatized) | splash10-0f79-0900000000-31e0a7f3a4c3fd20ce3c | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-EI-TOF (Non-derivatized) | splash10-0udr-0900000000-6d1ec3a7649c0e7a704b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-EI-TOF (Non-derivatized) | splash10-0f79-0900000000-468e3da761e4d86223b1 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-EI-TOF (Non-derivatized) | splash10-0fki-5900000000-0569fb6d8502d22d2b7e | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Hypotaurine GC-MS (Non-derivatized) | splash10-0f79-0900000000-457c0f4ff1563d09a208 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Hypotaurine GC-MS (Non-derivatized) - 70eV, Positive | splash10-000x-9000000000-ebfa02ce7482006f53d1 | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Hypotaurine GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-0a4i-2900000000-123846d2dc8e7cf97137 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-03di-9000000000-6b0caea8d77740b01ec0 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-03di-9000000000-a5ce3fbe0fd14222c3c8 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-03di-9000000000-32fdc560d98f6fd6be25 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative-QTOF | splash10-03di-9000000000-5d74861c832bbb4f97fc | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Negative-QTOF | splash10-03di-9300000000-85302e310b341400f079 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ , negative-QTOF | splash10-0a4i-2900000000-123846d2dc8e7cf97137 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ , negative-QTOF | splash10-03di-9000000000-6b0caea8d77740b01ec0 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ , negative-QTOF | splash10-03di-9000000000-a5ce3fbe0fd14222c3c8 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ , negative-QTOF | splash10-03di-9000000000-32fdc560d98f6fd6be25 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ , negative-QTOF | splash10-03di-9000000000-5d74861c832bbb4f97fc | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QTOF , negative-QTOF | splash10-03di-9300000000-85302e310b341400f079 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine , negative-QTOF | splash10-03di-9200000000-dcffc1a76d64346e7107 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-01ox-9800000000-f121cbb956b42fc5e20e | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-014i-9300000000-eec8c8d22a33d4e8da3b | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-014i-9200000000-f461253126e9b74c7d80 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-03di-0940100000-886905799c16b4d25a5f | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0006-9000000000-a34ee572b02492eaefef | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0006-9000000000-ff53f17dcba5f9b8c507 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-0900000000-a3ccc18b5af8fddfdffa | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-004i-0920000000-9c1032a24816a6221477 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0006-9000000000-6354cd263d6d293f1d61 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-0900000000-45a36787ef1f00274735 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-004i-0900000000-318c8a0e0325b2e98a06 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Hypotaurine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive-QTOF | splash10-03di-6900000000-26d55a4c69512ce8ae79 | 2012-08-31 | HMDB team, MONA | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-24 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|
| General References | - Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Dominy J, Eller S, Dawson R Jr: Building biosynthetic schools: reviewing compartmentation of CNS taurine synthesis. Neurochem Res. 2004 Jan;29(1):97-103. [PubMed:14992267 ]
- Fontana M, Pecci L, Dupre S, Cavallini D: Antioxidant properties of sulfinates: protective effect of hypotaurine on peroxynitrite-dependent damage. Neurochem Res. 2004 Jan;29(1):111-6. [PubMed:14992269 ]
- Pitari G, Dupre S, Spirito A, Antonini G, Amicarelli F: Hypotaurine protection on cell damage by singlet oxygen. Adv Exp Med Biol. 2000;483:157-62. [PubMed:11787593 ]
- Grafe F, Wohlrab W, Neubert RH, Brandsch M: Functional characterization of sodium- and chloride-dependent taurine transport in human keratinocytes. Eur J Pharm Biopharm. 2004 Mar;57(2):337-41. [PubMed:15018993 ]
- Krieg RC, Uihlein D, Murthum T, Endlicher E, Hausmann F, Messmann H, Knuechel R: Improving the use of 5-aminolevulinic acid (ALA)-induced protoporphyrin IX (PPIX) for the gastrointestinal tract by esterification--an in vitro study. Cell Mol Biol (Noisy-le-grand). 2002 Dec;48(8):917-23. [PubMed:12699251 ]
- Guerin P, Menezo Y: Hypotaurine and taurine in gamete and embryo environments: de novo synthesis via the cysteine sulfinic acid pathway in oviduct cells. Zygote. 1995 Nov;3(4):333-43. [PubMed:8730898 ]
- Masuoka N, Yao K, Kinuta M, Ohta J, Wakimoto M, Ubuka T: High-performance liquid chromatographic determination of taurine and hypotaurine using 3,5-dinitrobenzoyl chloride as derivatizing reagent. J Chromatogr B Biomed Appl. 1994 Oct 3;660(1):31-5. [PubMed:7858721 ]
- Guerin P, Guillaud J, Menezo Y: Hypotaurine in spermatozoa and genital secretions and its production by oviduct epithelial cells in vitro. Hum Reprod. 1995 Apr;10(4):866-72. [PubMed:7650134 ]
- Holmes RP, Goodman HO, Shihabi ZK, Jarow JP: The taurine and hypotaurine content of human semen. J Androl. 1992 May-Jun;13(3):289-92. [PubMed:1601750 ]
- Mahadevan MM, Trounson AO: Removal of the cumulus oophorus from the human oocyte for in vitro fertilization. Fertil Steril. 1985 Feb;43(2):263-7. [PubMed:3967784 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
|
|---|