| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Expected but not Quantified |
|---|
| Creation Date | 2006-05-22 14:17:39 UTC |
|---|
| Update Date | 2022-03-07 02:49:13 UTC |
|---|
| HMDB ID | HMDB0002159 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | 3a,7a-Dihydroxy-5b-cholestanoyl-CoA |
|---|
| Description | 3a,7a-Dihydroxy-5b-cholestanoyl-CoA belongs to the class of organic compounds known as 2,3,4-saturated fatty acyl coas. These are acyl-CoAs carrying a 2,3,4-saturated fatty acyl chain. Based on a literature review very few articles have been published on 3a,7a-Dihydroxy-5b-cholestanoyl-CoA. |
|---|
| Structure | [H][C@@]12C[C@H](O)CC[C@]1(C)C1CC[C@]3(C)C(CCC3C1[C@H](O)C2)C(C)CCCC(C)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C(N)N=CN=C12 InChI=1S/C48H80N7O19P3S/c1-26(30-10-11-31-36-32(13-16-48(30,31)6)47(5)15-12-29(56)20-28(47)21-33(36)57)8-7-9-27(2)45(62)78-19-18-50-35(58)14-17-51-43(61)40(60)46(3,4)23-71-77(68,69)74-76(66,67)70-22-34-39(73-75(63,64)65)38(59)44(72-34)55-25-54-37-41(49)52-24-53-42(37)55/h24-34,36,38-40,44,56-57,59-60H,7-23H2,1-6H3,(H,50,58)(H,51,61)(H,66,67)(H,68,69)(H2,49,52,53)(H2,63,64,65)/t26?,27?,28-,29+,30?,31?,32?,33+,34+,36?,38+,39+,40?,44+,47-,48+/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| 3-alpha,7-alpha-Dihydroxy-5-beta-cholestanoyl-CoA | HMDB | | 3-alpha,7-alpha-Dihydroxy-5-beta-cholestanoyl-coenzyme A | HMDB | | 3alpha,7alpha-Dihydroxy-5beta-cholestanoyl-CoA | HMDB | | 3alpha,7alpha-Dihydroxy-5beta-cholestanoyl-coenzyme A | HMDB | | DHCA | HMDB |
|
|---|
| Chemical Formula | C48H80N7O19P3S |
|---|
| Average Molecular Weight | 1184.171 |
|---|
| Monoisotopic Molecular Weight | 1183.444253639 |
|---|
| IUPAC Name | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-2-[({[({3-[(2-{[2-({6-[(2S,5R,7S,9R,15R)-5,9-dihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]-2-methylheptanoyl}sulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]-3-hydroxy-2,2-dimethylpropoxy}(hydroxy)phosphoryl)oxy](hydroxy)phosphoryl}oxy)methyl]-4-hydroxyoxolan-3-yl]oxy}phosphonic acid |
|---|
| Traditional Name | [(2R,3S,4R,5R)-5-(6-aminopurin-9-yl)-2-{[({3-[(2-{[2-({6-[(2S,5R,7S,9R,15R)-5,9-dihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]-2-methylheptanoyl}sulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]-3-hydroxy-2,2-dimethylpropoxy(hydroxy)phosphoryl}oxy(hydroxy)phosphoryl)oxy]methyl}-4-hydroxyoxolan-3-yl]oxyphosphonic acid |
|---|
| CAS Registry Number | 2461-62-3 |
|---|
| SMILES | [H][C@@]12C[C@H](O)CC[C@]1(C)C1CC[C@]3(C)C(CCC3C1[C@H](O)C2)C(C)CCCC(C)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C(N)N=CN=C12 |
|---|
| InChI Identifier | InChI=1S/C48H80N7O19P3S/c1-26(30-10-11-31-36-32(13-16-48(30,31)6)47(5)15-12-29(56)20-28(47)21-33(36)57)8-7-9-27(2)45(62)78-19-18-50-35(58)14-17-51-43(61)40(60)46(3,4)23-71-77(68,69)74-76(66,67)70-22-34-39(73-75(63,64)65)38(59)44(72-34)55-25-54-37-41(49)52-24-53-42(37)55/h24-34,36,38-40,44,56-57,59-60H,7-23H2,1-6H3,(H,50,58)(H,51,61)(H,66,67)(H,68,69)(H2,49,52,53)(H2,63,64,65)/t26?,27?,28-,29+,30?,31?,32?,33+,34+,36?,38+,39+,40?,44+,47-,48+/m0/s1 |
|---|
| InChI Key | SBYLHTNKEWSLBA-CPRJDYHCSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as 2,3,4-saturated fatty acyl coas. These are acyl-CoAs carrying a 2,3,4-saturated fatty acyl chain. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
| Direct Parent | 2,3,4-saturated fatty acyl CoAs |
|---|
| Alternative Parents | |
|---|
| Substituents | - Coenzyme a or derivatives
- Purine ribonucleoside 3',5'-bisphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside diphosphate
- Steroidal glycoside
- Dihydroxy bile acid, alcohol, or derivatives
- Hydroxy bile acid, alcohol, or derivatives
- Cholane-skeleton
- Bile acid, alcohol, or derivatives
- 3-hydroxysteroid
- Hydroxysteroid
- 3-alpha-hydroxysteroid
- 7-hydroxysteroid
- Steroid
- Ribonucleoside 3'-phosphate
- Pentose phosphate
- Pentose-5-phosphate
- Beta amino acid or derivatives
- Glycosyl compound
- N-glycosyl compound
- Organic pyrophosphate
- 6-aminopurine
- Pentose monosaccharide
- Monosaccharide phosphate
- Purine
- Imidazopyrimidine
- Monoalkyl phosphate
- Aminopyrimidine
- Pyrimidine
- Monosaccharide
- N-acyl-amine
- Phosphoric acid ester
- Fatty amide
- N-substituted imidazole
- Imidolactam
- Organic phosphoric acid derivative
- Alkyl phosphate
- Azole
- Tetrahydrofuran
- Cyclic alcohol
- Imidazole
- Heteroaromatic compound
- Amino acid or derivatives
- Carbothioic s-ester
- Carboxamide group
- Thiocarboxylic acid ester
- Secondary carboxylic acid amide
- Secondary alcohol
- Thiocarboxylic acid or derivatives
- Oxacycle
- Sulfenyl compound
- Azacycle
- Carboxylic acid derivative
- Organoheterocyclic compound
- Primary amine
- Alcohol
- Carbonyl group
- Organic oxygen compound
- Organosulfur compound
- Organooxygen compound
- Organopnictogen compound
- Amine
- Organic nitrogen compound
- Organonitrogen compound
- Hydrocarbon derivative
- Organic oxide
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesNot Available | Show more...
|---|
| Spectra |
|---|
| MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 10V, Positive-QTOF | splash10-000i-1900112000-0c75e0a93454d43ab753 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 20V, Positive-QTOF | splash10-000i-0900314000-ee39276da14ef1ec86d2 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 40V, Positive-QTOF | splash10-000i-1900000010-5f6513051536bb79e578 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 10V, Negative-QTOF | splash10-00lr-2901332400-00267947c2aa67bae2f7 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 20V, Negative-QTOF | splash10-003r-3900111010-35ffb32be4339618cff9 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 40V, Negative-QTOF | splash10-057i-6900100000-125c30cd4205b3d5be3d | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 10V, Positive-QTOF | splash10-0159-0900000000-eb50c2a058862834d250 | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 20V, Positive-QTOF | splash10-0159-0900102622-13f5492940fca24737d0 | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 40V, Positive-QTOF | splash10-004i-0000119000-c789136e01e3fc6660ea | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 10V, Negative-QTOF | splash10-001i-0900000000-787baca3a444ef57d1b3 | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 20V, Negative-QTOF | splash10-001i-2900301100-c831b98264288f5ecac5 | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestanoyl-CoA 40V, Negative-QTOF | splash10-0170-6605603900-a71bad576505bfdacb19 | 2021-09-25 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | - Extracellular
- Membrane
- Endoplasmic reticulum
- Peroxisome
|
|---|
| Biospecimen Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
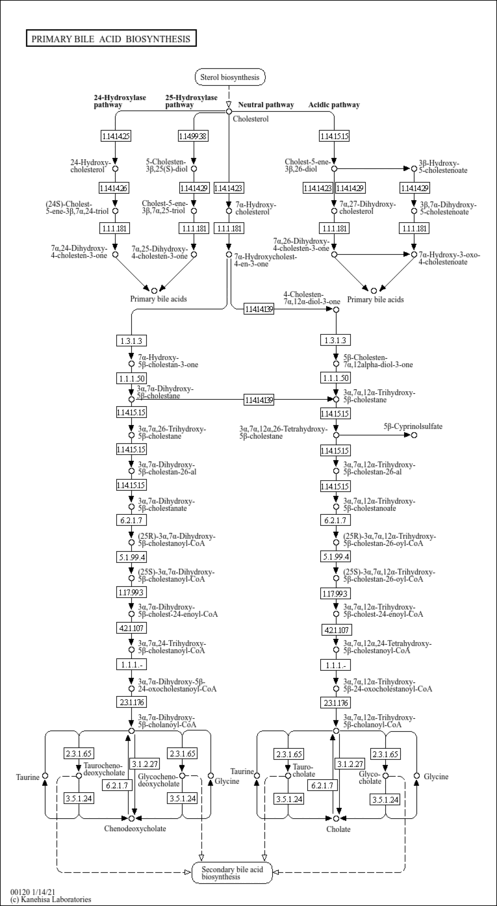
| Pathways | |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | None |
|---|
| Associated OMIM IDs | None |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB022875 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 24850079 |
|---|
| KEGG Compound ID | C04644 |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 440420 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | Not Available |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | DHCHOLESTANCOA |
|---|
| MarkerDB ID | Not Available |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Prydz K, Kase BF, Bjorkhem I, Pedersen JI: Subcellular localization of 3 alpha, 7 alpha-dihydroxy- and 3 alpha,7 alpha,12 alpha-trihydroxy-5 beta-cholestanoyl-coenzyme A ligase(s) in rat liver. J Lipid Res. 1988 Aug;29(8):997-1004. [PubMed:3183523 ]
|
|---|