| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2006-05-22 14:17:39 UTC |
|---|
| Update Date | 2023-02-21 17:16:13 UTC |
|---|
| HMDB ID | HMDB0002166 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | (S)-beta-Aminoisobutyric acid |
|---|
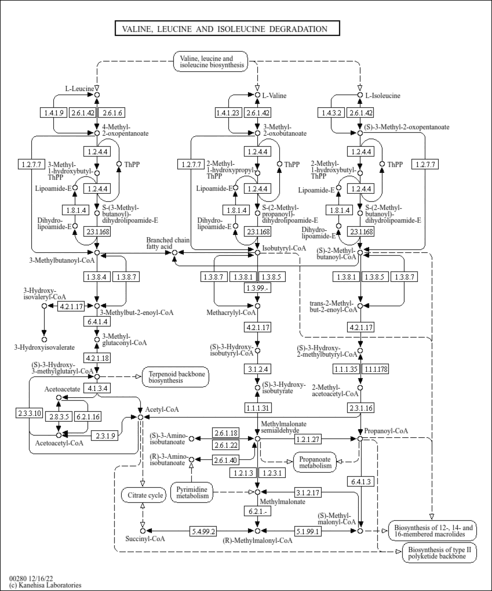
| Description | beta-Aminoisobutyric acid is a non-protein amino acid originating from the catabolism of thymine and valine. The concentration of beta-aminoisobutyric acid is normally low in urine as beta-aminoisobutyric acid is further catabolized by beta-aminoisobutyrate aminotransferases to methylmalonic acid semialdehyde and propionyl-CoA. beta-Aminoisobutyric acid occurs in two isomeric forms and both enantiomers of beta-aminoisobutyric acid can be detected in human urine and plasma. In plasma, the S-enantiomer is the predominant type due to active renal reabsorption. In contrast, urine almost exclusively contains the R-enantiomer of beta-aminoisobutyric acid, which is eliminated both by filtration and tubular secretion. Persistently increased levels of beta-aminoisobutyric acid have been observed in individuals with a deficiency of R (-)-beta-aminoisobutyrate-pyruvate aminotransferase. In addition, transient high levels of beta-aminoisobutyric acid have been observed under a variety of pathological conditions such as lead poisoning, starvation, in total body irradiation, and in a number of malignancies. The S-enantiomer of beta-aminoisobutyric acid is predominantly derived from the catabolism of valine. It has been suggested that altered homeostasis of beta-alanine underlies some of the clinical abnormalities encountered in patients with a dihydropyrimidine dehydrogenase (DPD) deficiency. DPD constitutes the first step of the pyrimidine degradation pathway, in which the pyrimidine bases uracil and thymine are catabolized to beta-alanine and the R-enantiomer of beta-aminoisobutyric acid respectively. In normal individuals with an intact pyrimidine degradation pathway, R-methylmalonic acid semialdehyde can be synthesized directly from the catabolism of thymine. Hence, there might be less cross-over between the valine and thymine pathway, allowing the conversion of S-methylmalonic acid semialdehyde into S-beta-aminoisobutyric acid and the subsequent accumulation of S-beta-aminoisobutyric acid in plasma (PMID: 14705962 , 14292857 , 14453202 ). |
|---|
| Structure | InChI=1S/C4H9NO2/c1-3(2-5)4(6)7/h3H,2,5H2,1H3,(H,6,7)/t3-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (S)-3-Amino-2-methylpropanoic acid | ChEBI | | (S)-3-Amino-isobutanoic acid | ChEBI | | (S)-3-Amino-isobutyric acid | ChEBI | | L-3-Amino-isobutanoic acid | ChEBI | | L-3-Amino-isobutyric acid | ChEBI | | (S)-3-Aminoisobutyrate | Kegg | | L-3-Aminoisobutyrate | Kegg | | (S)-3-Aminoisobutanoate | Kegg | | (S)-3-Amino-2-methylpropanoate | Kegg | | (S)-beta-Aminoisobutyrate | Kegg | | (S)-3-Amino-isobutanoate | Generator | | (S)-3-Amino-isobutyrate | Generator | | L-3-Amino-isobutanoate | Generator | | L-3-Amino-isobutyrate | Generator | | (S)-3-Aminoisobutyric acid | Generator | | L-3-Aminoisobutyric acid | Generator | | (S)-3-Aminoisobutanoic acid | Generator | | (S)-b-Aminoisobutyrate | Generator | | (S)-b-Aminoisobutyric acid | Generator | | (S)-Β-aminoisobutyrate | Generator | | (S)-Β-aminoisobutyric acid | Generator | | (S)-beta-Aminoisobutyric acid | ChEBI | | (+)-a-Methyl-b-alanine | HMDB | | (+)-alpha-Methyl-beta-alanine | HMDB | | (+)-b-Aminoisobutyric acid | HMDB | | (+)-beta-Aminoisobutyric acid | HMDB | | (S)-3-amino-2-Methyl-propanoate | HMDB | | (S)-3-amino-2-Methyl-propanoic acid | HMDB | | L-2-Methyl-b-alanine | HMDB | | L-2-Methyl-beta-alanine | HMDB | | L-3-amino-2-Methylpropanoate | HMDB | | L-3-amino-2-Methylpropanoic acid | HMDB | | L-3-amino-2-Methylpropionic acid | HMDB | | L-b-Aminoisobutyrate | HMDB | | L-b-Aminoisobutyric acid | HMDB | | L-beta-Aminoisobutyrate | HMDB | | L-beta-Aminoisobutyric acid | HMDB | | S-b-Aminoisobutyrate | HMDB | | S-beta-Aminoisobutyrate | HMDB | | S-beta-Aminoisobutyric acid | HMDB |
|
|---|
| Chemical Formula | C4H9NO2 |
|---|
| Average Molecular Weight | 103.1198 |
|---|
| Monoisotopic Molecular Weight | 103.063328537 |
|---|
| IUPAC Name | (2S)-3-amino-2-methylpropanoic acid |
|---|
| Traditional Name | L-β-aminoisobutyric acid |
|---|
| CAS Registry Number | 4249-19-8 |
|---|
| SMILES | C[C@@H](CN)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C4H9NO2/c1-3(2-5)4(6)7/h3H,2,5H2,1H3,(H,6,7)/t3-/m0/s1 |
|---|
| InChI Key | QCHPKSFMDHPSNR-VKHMYHEASA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as beta amino acids and derivatives. These are amino acids having a (-NH2) group attached to the beta carbon atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Beta amino acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Beta amino acid or derivatives
- Amino acid
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Amine
- Organic oxide
- Hydrocarbon derivative
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Organopnictogen compound
- Primary aliphatic amine
- Organic oxygen compound
- Carbonyl group
- Organic nitrogen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 175 - 177 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 1.28 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 8.5725 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 6.16 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 350.2 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 484.1 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 326.7 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 51.6 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 211.6 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 67.3 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 275.9 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 229.6 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 743.1 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 557.3 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 35.6 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 620.6 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 199.4 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 269.0 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 636.1 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 471.1 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 339.3 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| (S)-beta-Aminoisobutyric acid,1TMS,isomer #1 | C[C@@H](CN)C(=O)O[Si](C)(C)C | 1051.8 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,1TMS,isomer #2 | C[C@@H](CN[Si](C)(C)C)C(=O)O | 1206.4 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TMS,isomer #1 | C[C@@H](CN[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1230.4 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TMS,isomer #1 | C[C@@H](CN[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1249.7 | Standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TMS,isomer #1 | C[C@@H](CN[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1331.9 | Standard polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TMS,isomer #2 | C[C@@H](CN([Si](C)(C)C)[Si](C)(C)C)C(=O)O | 1412.2 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TMS,isomer #2 | C[C@@H](CN([Si](C)(C)C)[Si](C)(C)C)C(=O)O | 1357.2 | Standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TMS,isomer #2 | C[C@@H](CN([Si](C)(C)C)[Si](C)(C)C)C(=O)O | 1608.9 | Standard polar | 33892256 | | (S)-beta-Aminoisobutyric acid,3TMS,isomer #1 | C[C@@H](CN([Si](C)(C)C)[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1464.4 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,3TMS,isomer #1 | C[C@@H](CN([Si](C)(C)C)[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1414.9 | Standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,3TMS,isomer #1 | C[C@@H](CN([Si](C)(C)C)[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1339.4 | Standard polar | 33892256 | | (S)-beta-Aminoisobutyric acid,1TBDMS,isomer #1 | C[C@@H](CN)C(=O)O[Si](C)(C)C(C)(C)C | 1282.3 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,1TBDMS,isomer #2 | C[C@@H](CN[Si](C)(C)C(C)(C)C)C(=O)O | 1450.9 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TBDMS,isomer #1 | C[C@@H](CN[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1668.6 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TBDMS,isomer #1 | C[C@@H](CN[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1679.3 | Standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TBDMS,isomer #1 | C[C@@H](CN[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1627.1 | Standard polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TBDMS,isomer #2 | C[C@@H](CN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O | 1828.1 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TBDMS,isomer #2 | C[C@@H](CN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O | 1761.4 | Standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,2TBDMS,isomer #2 | C[C@@H](CN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O | 1785.6 | Standard polar | 33892256 | | (S)-beta-Aminoisobutyric acid,3TBDMS,isomer #1 | C[C@@H](CN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2095.5 | Semi standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,3TBDMS,isomer #1 | C[C@@H](CN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2054.2 | Standard non polar | 33892256 | | (S)-beta-Aminoisobutyric acid,3TBDMS,isomer #1 | C[C@@H](CN([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1764.4 | Standard polar | 33892256 |
|
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - (S)-beta-Aminoisobutyric acid GC-MS (Non-derivatized) - 70eV, Positive | splash10-001i-9000000000-59e52601f90d8e636d61 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - (S)-beta-Aminoisobutyric acid GC-MS (1 TMS) - 70eV, Positive | splash10-00ar-9200000000-0eb92539979a9fa36dbd | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - (S)-beta-Aminoisobutyric acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 1V, negative-QTOF | splash10-0udi-1900000000-aaccebe5d4a868bc32bf | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 2V, negative-QTOF | splash10-0udi-2900000000-4f4e49fe169d5c21efc1 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 2V, negative-QTOF | splash10-0udi-4900000000-f32e82420f5efb60456e | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 3V, negative-QTOF | splash10-0uk9-6900000000-cb0ef82b397e24372142 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 3V, negative-QTOF | splash10-0uk9-8900000000-31fc31847c8771693d15 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 3V, negative-QTOF | splash10-0fk9-9600000000-6379643466e30660a05b | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 4V, negative-QTOF | splash10-00di-9300000000-97e8a7125780e41fceaf | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 5V, negative-QTOF | splash10-05fr-9200000000-8f0488fe62603d121536 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 5V, negative-QTOF | splash10-0ab9-9100000000-84e07fbee3e332502265 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 6V, negative-QTOF | splash10-0a4i-9000000000-3634b730902e2e080b56 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid n/a 7V, negative-QTOF | splash10-00di-9000000000-1b9078e3902a11473fa0 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 0V, positive-QTOF | splash10-0udi-1900000000-76d19f44560e6b1b6f80 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 0V, positive-QTOF | splash10-0udi-1900000000-bd4e85f35b448a2b7540 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 0V, positive-QTOF | splash10-0udi-2900000000-b1371ec53be10ec9c411 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 1V, positive-QTOF | splash10-0udi-3900000000-d16b3475fcdfed5038d5 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 1V, positive-QTOF | splash10-0udi-4900000000-cd34930bbcb805c7525f | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 1V, positive-QTOF | splash10-0udr-5900000000-4b73ac1046680185a981 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 1V, positive-QTOF | splash10-0udr-6900000000-a911da27982a560a6cf1 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid Orbitrap 1V, positive-QTOF | splash10-0udr-8900000000-91a255aae27e4e4aabb8 | 2020-07-22 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid 10V, Positive-QTOF | splash10-0f79-9300000000-30669ed04aadf3d3f040 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid 20V, Positive-QTOF | splash10-000l-9000000000-ca65637476e64152c44e | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid 40V, Positive-QTOF | splash10-0006-9000000000-04d1648391d3f90db226 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid 10V, Negative-QTOF | splash10-0udi-3900000000-ec54fcce079024a62344 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid 20V, Negative-QTOF | splash10-0zfr-9500000000-0394f399cdf5df93c5df | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - (S)-beta-Aminoisobutyric acid 40V, Negative-QTOF | splash10-0a4l-9000000000-d0c495c908ed905558a2 | 2015-09-15 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|