| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected but not Quantified |
|---|
| Creation Date | 2008-08-12 13:40:35 UTC |
|---|
| Update Date | 2022-09-22 18:34:19 UTC |
|---|
| HMDB ID | HMDB0006809 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Nicotinic acid ribonucleoside |
|---|
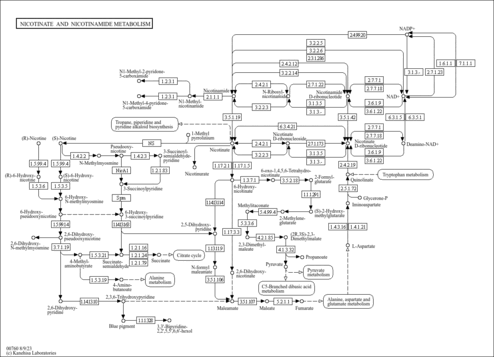
| Description | Nicotinate D-ribonucleoside, also known as nicotinic acid riboside or beta-D-ribosylnicotinate, belongs to the class of organic compounds known as glycosylamines. Glycosylamines are compounds consisting of an amine with a beta-N-glycosidic bond to a carbohydrate, thus forming a cyclic hemiaminal ether bond (alpha-amino ether). Nicotinate D-ribonucleoside is an extremely weak basic (essentially neutral) compound (based on its pKa). Nicotinate D-ribonucleoside exists in all living species, ranging from bacteria to humans. Within humans, nicotinate D-ribonucleoside participates in a number of enzymatic reactions. In particular, nicotinate D-ribonucleoside can be converted into nicotinic acid mononucleotide; which is catalyzed by the enzyme nicotinamide riboside kinase 1. In addition, nicotinate D-ribonucleoside can be biosynthesized from nicotinic acid mononucleotide; which is mediated by the enzyme cytosolic purine 5'-nucleotidase. A pyridine nucleoside consisting of nicotinic acid with a beta-D-ribofuranosyl moiety at the 1-position. In humans, nicotinate D-ribonucleoside is involved in nicotinate and nicotinamide metabolism. |
|---|
| Structure | OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)[N+]1=CC=CC(=C1)C(O)=O InChI=1S/C11H13NO6/c13-5-7-8(14)9(15)10(18-7)12-3-1-2-6(4-12)11(16)17/h1-4,7-10,13-15H,5H2/p+1/t7-,8-,9-,10-/m1/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| Nicotinic acid riboside | ChEBI | | beta-D-Ribosylnicotinate | Kegg | | Nicotinate riboside | Generator | | b-D-Ribosylnicotinate | Generator | | b-D-Ribosylnicotinic acid | Generator | | beta-D-Ribosylnicotinic acid | Generator | | Β-D-ribosylnicotinate | Generator | | Β-D-ribosylnicotinic acid | Generator | | Nicotinic acid D-ribonucleoside | Generator | | D-Ribosylnicotinate | HMDB | | Nicotinic acid ribose | HMDB | | Ribosylnicotinate | HMDB | | Nicotinate ribonucleoside | HMDB | | Nicotinate ribose | HMDB | | Nicotinic acid ribonucleoside | HMDB | | 3-Carboxy-1-beta-D-ribofuranosylpyridinium | HMDB | | 3-Carboxy-1-β-D-ribofuranosylpyridinium | HMDB | | Nicotinic riboside | HMDB |
| Show more...
|---|
| Chemical Formula | C11H14NO6 |
|---|
| Average Molecular Weight | 256.232 |
|---|
| Monoisotopic Molecular Weight | 256.082112185 |
|---|
| IUPAC Name | 3-carboxy-1-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-1lambda5-pyridin-1-ylium |
|---|
| Traditional Name | 3-carboxy-1-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-1lambda5-pyridin-1-ylium |
|---|
| CAS Registry Number | 4013-06-3 |
|---|
| SMILES | OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)[N+]1=CC=CC(=C1)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C11H13NO6/c13-5-7-8(14)9(15)10(18-7)12-3-1-2-6(4-12)11(16)17/h1-4,7-10,13-15H,5H2/p+1/t7-,8-,9-,10-/m1/s1 |
|---|
| InChI Key | PUEDDPCUCPRQNY-ZYUZMQFOSA-O |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as glycosylamines. Glycosylamines are compounds consisting of an amine with a beta-N-glycosidic bond to a carbohydrate, thus forming a cyclic hemiaminal ether bond (alpha-amino ether). |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Glycosylamines |
|---|
| Alternative Parents | |
|---|
| Substituents | - N-glycosyl compound
- Pentose monosaccharide
- Pyridine carboxylic acid
- Pyridine carboxylic acid or derivatives
- Monosaccharide
- Pyridine
- Pyridinium
- Vinylogous amide
- Heteroaromatic compound
- Tetrahydrofuran
- Secondary alcohol
- Organoheterocyclic compound
- Carboxylic acid derivative
- Oxacycle
- Azacycle
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organonitrogen compound
- Alcohol
- Primary alcohol
- Organic nitrogen compound
- Organic cation
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Nicotinic acid ribonucleoside,1TMS,isomer #1 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O)[C@@H]1O | 2435.3 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TMS,isomer #2 | C[Si](C)(C)O[C@@H]1[C@H](O)[C@@H](CO)O[C@H]1[N+]1=CC=CC(C(=O)O)=C1 | 2439.4 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TMS,isomer #3 | C[Si](C)(C)O[C@@H]1[C@@H](CO)O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@@H]1O | 2438.3 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TMS,isomer #4 | C[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O)[C@H]2O)=C1 | 2354.2 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TMS,isomer #1 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C)=C2)[C@H](O)[C@@H]1O | 2378.6 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TMS,isomer #2 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O[Si](C)(C)C)[C@@H]1O | 2410.2 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TMS,isomer #3 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O)[C@@H]1O[Si](C)(C)C | 2405.5 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TMS,isomer #4 | C[Si](C)(C)O[C@@H]1[C@@H](CO)O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@@H]1O[Si](C)(C)C | 2399.9 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TMS,isomer #5 | C[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O)[C@H]2O[Si](C)(C)C)=C1 | 2371.6 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TMS,isomer #6 | C[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O[Si](C)(C)C)[C@H]2O)=C1 | 2370.7 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TMS,isomer #1 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C)=C2)[C@H](O[Si](C)(C)C)[C@@H]1O | 2402.6 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TMS,isomer #2 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C)=C2)[C@H](O)[C@@H]1O[Si](C)(C)C | 2397.8 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TMS,isomer #3 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O[Si](C)(C)C)[C@@H]1O[Si](C)(C)C | 2390.1 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TMS,isomer #4 | C[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O[Si](C)(C)C)[C@H]2O[Si](C)(C)C)=C1 | 2385.2 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,4TMS,isomer #1 | C[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C)=C2)[C@H](O[Si](C)(C)C)[C@@H]1O[Si](C)(C)C | 2414.9 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O)[C@@H]1O | 2686.1 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)O[C@@H]1[C@H](O)[C@@H](CO)O[C@H]1[N+]1=CC=CC(C(=O)O)=C1 | 2674.6 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)O[C@@H]1[C@@H](CO)O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@@H]1O | 2686.3 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,1TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O)[C@H]2O)=C1 | 2622.8 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C(C)(C)C)=C2)[C@H](O)[C@@H]1O | 2879.2 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O[Si](C)(C)C(C)(C)C)[C@@H]1O | 2911.9 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O)[C@@H]1O[Si](C)(C)C(C)(C)C | 2910.9 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)O[C@@H]1[C@@H](CO)O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@@H]1O[Si](C)(C)C(C)(C)C | 2893.9 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TBDMS,isomer #5 | CC(C)(C)[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O)[C@H]2O[Si](C)(C)C(C)(C)C)=C1 | 2881.1 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,2TBDMS,isomer #6 | CC(C)(C)[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O[Si](C)(C)C(C)(C)C)[C@H]2O)=C1 | 2898.5 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C(C)(C)C)=C2)[C@H](O[Si](C)(C)C(C)(C)C)[C@@H]1O | 3107.8 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C(C)(C)C)=C2)[C@H](O)[C@@H]1O[Si](C)(C)C(C)(C)C | 3103.4 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O)=C2)[C@H](O[Si](C)(C)C(C)(C)C)[C@@H]1O[Si](C)(C)C(C)(C)C | 3115.7 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,3TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)OC(=O)C1=CC=C[N+]([C@@H]2O[C@H](CO)[C@@H](O[Si](C)(C)C(C)(C)C)[C@H]2O[Si](C)(C)C(C)(C)C)=C1 | 3101.0 | Semi standard non polar | 33892256 | | Nicotinic acid ribonucleoside,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC[C@H]1O[C@@H]([N+]2=CC=CC(C(=O)O[Si](C)(C)C(C)(C)C)=C2)[C@H](O[Si](C)(C)C(C)(C)C)[C@@H]1O[Si](C)(C)C(C)(C)C | 3262.7 | Semi standard non polar | 33892256 |
| Show more...
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - Nicotinic acid ribonucleoside GC-MS (Non-derivatized) - 70eV, Positive | splash10-009f-9640000000-9ecd90015c00728b3b07 | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Nicotinic acid ribonucleoside GC-MS (4 TMS) - 70eV, Positive | splash10-0fa9-9442740000-0573803f81575b22f5c8 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Nicotinic acid ribonucleoside GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Nicotinic acid ribonucleoside GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 10V, Negative-QTOF | splash10-0a4i-0190000000-c2d0a64c0eb3d4ae2bfa | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 20V, Negative-QTOF | splash10-0a4i-0490000000-bb44dd196c6e9a647df7 | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 40V, Negative-QTOF | splash10-0zgu-9600000000-d17b549da8f5e8b2dd2b | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 10V, Positive-QTOF | splash10-0a4i-0090000000-384ec325284b36db888b | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 20V, Positive-QTOF | splash10-002b-2190000000-5ab972cf9d81893b8388 | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 40V, Positive-QTOF | splash10-0006-9510000000-8e2f234fa7933273a714 | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 10V, Positive-QTOF | splash10-06e9-1950000000-ae12a3b253bcd6d48ccc | 2021-09-23 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 20V, Positive-QTOF | splash10-001i-9510000000-cb2cf76989881501793c | 2021-09-23 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Nicotinic acid ribonucleoside 40V, Positive-QTOF | splash10-001i-9100000000-9d7edc1ec5f0c17e5711 | 2021-09-23 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | Not Available |
|---|
| Biospecimen Locations | |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Not Available | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details |
|
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Expected but not Quantified | Not Quantified | Not Specified | Not Specified | Cancer patients undergoing total body irradiation | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Urine | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Cancer patients undergoing total body irradiation | | details |
|
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Colorectal cancer |
|---|
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB024094 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 141636 |
|---|
| KEGG Compound ID | C05841 |
|---|
| BioCyc ID | CPD-8259 |
|---|
| BiGG ID | 46613 |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 161234 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 27748 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | NICRNS |
|---|
| MarkerDB ID | Not Available |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | Not Available |
|---|