| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Expected but not Quantified |
|---|
| Creation Date | 2008-08-14 18:18:11 UTC |
|---|
| Update Date | 2022-03-07 02:49:34 UTC |
|---|
| HMDB ID | HMDB0006895 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
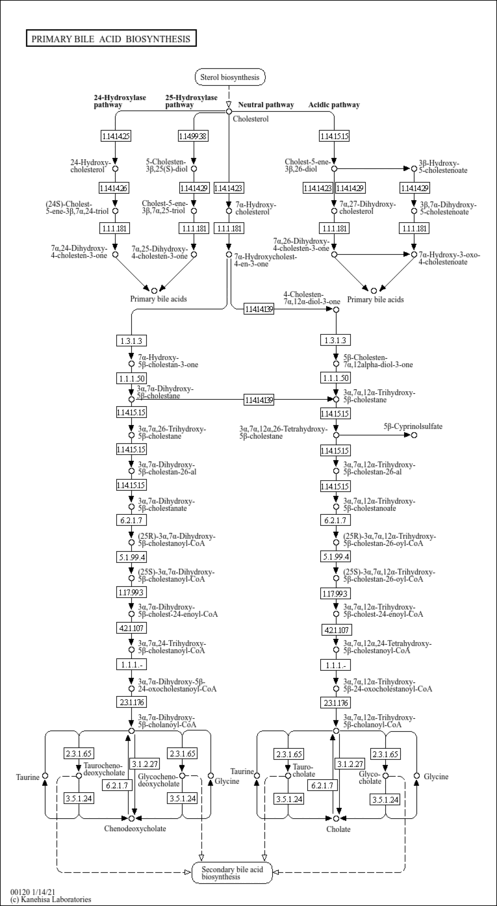
| Common Name | 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA |
|---|
| Description | 3alpha,7alpha-Dihydroxy-5beta-cholest-24-enoyl-CoA is an intermediate involved in bile acid synthesis, specifically in the synthesis of chenodeoxyglycocholate. Bile acids are steroid acids found predominantly in the bile of mammals. The distinction between different bile acids is minute, depending only on the presence or absence of hydroxyl groups on positions 3, 7, and 12. Bile acids are physiological detergents that facilitate excretion, absorption, and transport of fats and sterols in the intestine and liver. Bile acids are also steroidal amphipathic molecules derived from the catabolism of cholesterol. They modulate bile flow and lipid secretion, are essential for the absorption of dietary fats and vitamins, and have been implicated in the regulation of all the key enzymes involved in cholesterol homeostasis. Bile acids recirculate through the liver, bile ducts, small intestine and portal vein to form an enterohepatic circuit. They exist as anions at physiological pH and, consequently, require a carrier for transport across the membranes of the enterohepatic tissues. The unique detergent properties of bile acids are essential for the digestion and intestinal absorption of hydrophobic nutrients. Bile acids have potent toxic properties (e.g. membrane disruption) and there are a plethora of mechanisms to limit their accumulation in blood and tissues (PMID: 11316487 , 16037564 , 12576301 , 11907135 ). |
|---|
| Structure | CC(CC\C=C(/C)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C1N=CN=C2N)C1CCC2C3[C@H](O)C[C@]4([H])C[C@H](O)CC[C@]4(C)C3CC[C@]12C InChI=1S/C48H78N7O19P3S/c1-26(30-10-11-31-36-32(13-16-48(30,31)6)47(5)15-12-29(56)20-28(47)21-33(36)57)8-7-9-27(2)45(62)78-19-18-50-35(58)14-17-51-43(61)40(60)46(3,4)23-71-77(68,69)74-76(66,67)70-22-34-39(73-75(63,64)65)38(59)44(72-34)55-25-54-37-41(49)52-24-53-42(37)55/h9,24-26,28-34,36,38-40,44,56-57,59-60H,7-8,10-23H2,1-6H3,(H,50,58)(H,51,61)(H,66,67)(H,68,69)(H2,49,52,53)(H2,63,64,65)/b27-9+/t26?,28-,29+,30?,31?,32?,33+,34+,36?,38+,39+,40?,44+,47-,48+/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| 3a,7a-Dihydroxy-5b-cholest-24-enoyl-coenzyme A | HMDB | | 3alpha,7alpha-Dihydroxy-5beta-cholest-24-enoyl-CoA | HMDB | | 3alpha,7alpha-Dihydroxy-5beta-cholest-24-enoyl-coenzyme A | HMDB |
|
|---|
| Chemical Formula | C48H78N7O19P3S |
|---|
| Average Molecular Weight | 1182.155 |
|---|
| Monoisotopic Molecular Weight | 1181.428603575 |
|---|
| IUPAC Name | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-2-({[({[3-({2-[(2-{[(2E)-6-[(2S,5R,7S,9R,15R)-5,9-dihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]-2-methylhept-2-enoyl]sulfanyl}ethyl)carbamoyl]ethyl}carbamoyl)-3-hydroxy-2,2-dimethylpropoxy](hydroxy)phosphoryl}oxy)(hydroxy)phosphoryl]oxy}methyl)-4-hydroxyoxolan-3-yl]oxy}phosphonic acid |
|---|
| Traditional Name | [(2R,3S,4R,5R)-5-(6-aminopurin-9-yl)-2-[({[3-({2-[(2-{[(2E)-6-[(2S,5R,7S,9R,15R)-5,9-dihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]-2-methylhept-2-enoyl]sulfanyl}ethyl)carbamoyl]ethyl}carbamoyl)-3-hydroxy-2,2-dimethylpropoxy(hydroxy)phosphoryl]oxy(hydroxy)phosphoryl}oxy)methyl]-4-hydroxyoxolan-3-yl]oxyphosphonic acid |
|---|
| CAS Registry Number | Not Available |
|---|
| SMILES | CC(CC\C=C(/C)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C1N=CN=C2N)C1CCC2C3[C@H](O)C[C@]4([H])C[C@H](O)CC[C@]4(C)C3CC[C@]12C |
|---|
| InChI Identifier | InChI=1S/C48H78N7O19P3S/c1-26(30-10-11-31-36-32(13-16-48(30,31)6)47(5)15-12-29(56)20-28(47)21-33(36)57)8-7-9-27(2)45(62)78-19-18-50-35(58)14-17-51-43(61)40(60)46(3,4)23-71-77(68,69)74-76(66,67)70-22-34-39(73-75(63,64)65)38(59)44(72-34)55-25-54-37-41(49)52-24-53-42(37)55/h9,24-26,28-34,36,38-40,44,56-57,59-60H,7-8,10-23H2,1-6H3,(H,50,58)(H,51,61)(H,66,67)(H,68,69)(H2,49,52,53)(H2,63,64,65)/b27-9+/t26?,28-,29+,30?,31?,32?,33+,34+,36?,38+,39+,40?,44+,47-,48+/m0/s1 |
|---|
| InChI Key | SEBZZAWTQNNGPK-YOKOLYSVSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as acyl coas. These are organic compounds containing a coenzyme A substructure linked to an acyl chain. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
| Direct Parent | Acyl CoAs |
|---|
| Alternative Parents | |
|---|
| Substituents | - Coenzyme a or derivatives
- Purine ribonucleoside 3',5'-bisphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside diphosphate
- Steroidal glycoside
- Dihydroxy bile acid, alcohol, or derivatives
- Hydroxy bile acid, alcohol, or derivatives
- Cholane-skeleton
- Bile acid, alcohol, or derivatives
- 3-hydroxysteroid
- Hydroxysteroid
- 3-alpha-hydroxysteroid
- 7-hydroxysteroid
- Steroid
- Ribonucleoside 3'-phosphate
- Pentose phosphate
- Pentose-5-phosphate
- Beta amino acid or derivatives
- Glycosyl compound
- N-glycosyl compound
- Organic pyrophosphate
- 6-aminopurine
- Pentose monosaccharide
- Monosaccharide phosphate
- Purine
- Imidazopyrimidine
- Monoalkyl phosphate
- Aminopyrimidine
- Pyrimidine
- Monosaccharide
- N-acyl-amine
- Phosphoric acid ester
- Fatty amide
- N-substituted imidazole
- Imidolactam
- Organic phosphoric acid derivative
- Alkyl phosphate
- Azole
- Tetrahydrofuran
- Cyclic alcohol
- Imidazole
- Heteroaromatic compound
- Amino acid or derivatives
- Carbothioic s-ester
- Carboxamide group
- Thiocarboxylic acid ester
- Secondary carboxylic acid amide
- Secondary alcohol
- Thiocarboxylic acid or derivatives
- Oxacycle
- Sulfenyl compound
- Azacycle
- Carboxylic acid derivative
- Organoheterocyclic compound
- Primary amine
- Alcohol
- Carbonyl group
- Organic oxygen compound
- Organosulfur compound
- Organooxygen compound
- Organopnictogen compound
- Amine
- Organic nitrogen compound
- Organonitrogen compound
- Hydrocarbon derivative
- Organic oxide
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesNot Available | Show more...
|---|
| Spectra |
|---|
| MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 10V, Positive-QTOF | splash10-000i-1900112000-3614385f2311d2073e3e | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 20V, Positive-QTOF | splash10-000i-0900315000-797cccaf3753018d956f | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 40V, Positive-QTOF | splash10-000i-1900001000-61457f78771a3bf83e48 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 10V, Negative-QTOF | splash10-01q9-2901331400-4cfedec67e85eb7be44d | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 20V, Negative-QTOF | splash10-001i-3901121010-afd20923316bb8e01988 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 40V, Negative-QTOF | splash10-057i-5900100000-37ffc9114d81e845dfb6 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 10V, Negative-QTOF | splash10-001i-0900000000-666ad42e52868013faf0 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 20V, Negative-QTOF | splash10-01sr-3600902200-f9e74c1c38fcce635049 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 40V, Negative-QTOF | splash10-014r-5204604900-8df87961a4c0ed8e78f2 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 10V, Positive-QTOF | splash10-001i-0900000110-84949aea2a7fdac5d446 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 20V, Positive-QTOF | splash10-000i-0900001310-f616923d144fd5c5bd08 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholest-24-enoyl-CoA 40V, Positive-QTOF | splash10-004i-0000119000-c7ffdca0daf17da9409f | 2021-09-24 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-25 | Wishart Lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | |
|---|
| Biospecimen Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | None |
|---|
| Associated OMIM IDs | None |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB024142 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 4444354 |
|---|
| KEGG Compound ID | C05447 |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | 45830 |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 5280796 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 27393 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | DHCHOLOYLCOA |
|---|
| MarkerDB ID | Not Available |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - St-Pierre MV, Kullak-Ublick GA, Hagenbuch B, Meier PJ: Transport of bile acids in hepatic and non-hepatic tissues. J Exp Biol. 2001 May;204(Pt 10):1673-86. [PubMed:11316487 ]
- Claudel T, Staels B, Kuipers F: The Farnesoid X receptor: a molecular link between bile acid and lipid and glucose metabolism. Arterioscler Thromb Vasc Biol. 2005 Oct;25(10):2020-30. Epub 2005 Jul 21. [PubMed:16037564 ]
- Chiang JY: Bile acid regulation of hepatic physiology: III. Bile acids and nuclear receptors. Am J Physiol Gastrointest Liver Physiol. 2003 Mar;284(3):G349-56. [PubMed:12576301 ]
- Davis RA, Miyake JH, Hui TY, Spann NJ: Regulation of cholesterol-7alpha-hydroxylase: BAREly missing a SHP. J Lipid Res. 2002 Apr;43(4):533-43. [PubMed:11907135 ]
|
|---|