| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2025-05-29 19:33:02 UTC |
|---|
| HMDB ID | HMDB0000158 |
|---|
| Secondary Accession Numbers | - HMDB0000647
- HMDB00158
- HMDB00647
|
|---|
| Metabolite Identification |
|---|
| Common Name | L-Tyrosine |
|---|
| Description | Tyrosine (Tyr) or L-tyrosine is an alpha-amino acid. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). Amino acids are organic compounds that contain amino (-NH2) and carboxyl (-COOH) functional groups, along with a side chain (R group) specific to each amino acid. L-tyrosine is one of 20 proteinogenic amino acids, i.e., the amino acids used in the biosynthesis of proteins. Tyrosine is found in all organisms ranging from bacteria to plants to animals. It is classified as a non-polar, uncharged (at physiological pH) aromatic amino acid. Tyrosine is a non-essential amino acid, meaning the body can synthesize it - usually from phenylalanine. The conversion of phenylalanine to tyrosine is catalyzed by the enzyme phenylalanine hydroxylase, a monooxygenase. This enzyme catalyzes the reaction causing the addition of a hydroxyl group to the end of the 6-carbon aromatic ring of phenylalanine, such that it becomes tyrosine. Tyrosine is found in many high-protein food products such as chicken, turkey, fish, milk, yogurt, cottage cheese, cheese, peanuts, almonds, pumpkin seeds, sesame seeds, soy products, lima beans, avocados and bananas. Tyrosine is one of the few amino acids that readily passes the blood-brain barrier. Once in the brain, it is a precursor for the neurotransmitters dopamine, norepinephrine and epinephrine, better known as adrenalin. These neurotransmitters are an important part of the body's sympathetic nervous system, and their concentrations in the body and brain are directly dependent upon dietary tyrosine. Tyrosine is not found in large concentrations throughout the body, probably because it is rapidly metabolized. Folic acid, copper and vitamin C are cofactor nutrients of these reactions. Tyrosine is also the precursor for hormones, including thyroid hormones (diiodotyrosine), catecholestrogens and the major human pigment, melanin. Tyrosine is an important amino acid in many proteins, peptides and even enkephalins, the body's natural pain reliever. Valine and other branched amino acids, and possibly tryptophan and phenylalanine may reduce tyrosine absorption. A number of genetic errors of tyrosine metabolism have been identified, such as hawkinsinuria and tyrosinemia I. The most common feature of these diseases is the increased amount of tyrosine in the blood, which is marked by decreased motor activity, lethargy and poor feeding. Infection and intellectual deficits may occur. Vitamin C supplements can help reverse these disease symptoms. High tyrosine concentrations have also been detected in septic patients (PMID: 99098 ; PMID: 27501420 ). This may reflect the breakdown of muscle tissues (leading to amino acid release) and the body’s differential metabolic capacity for different amino acids. Muscle tissue is easily able to oxidize branched chain amino acids to support its own energy requirements. Muscles are also able to metabolize alanine, glycine, proline, aspartate, glutamate, histidine, glutamine and serine for gluconeogenesis, but aromatic amino acids such as phenylalanine and tyrosine as well as many cysteine-containing amino acids are not as easily metabolized. This may account for the increase in the levels of tyrosine seen during sepsis (PMID: 99098 ). Independent of the occurrence of infection or injury, some adults can develop elevated tyrosine levels in their blood. This typically indicates a need for more vitamin C. More tyrosine is needed under stress, and tyrosine supplements prevent the stress-induced depletion of norepinephrine and can help alleviate biochemical depression. However, tyrosine may not be good for treating psychosis. Many antipsychotic medications apparently function by inhibiting tyrosine metabolism. L-Dopa, which is directly used in Parkinson's, is made from tyrosine. Tyrosine, the nutrient, can be used as an adjunct in the treatment of Parkinson's. Peripheral metabolism of tyrosine necessitates large doses of tyrosine, however, compared to L-Dopa (http://www.dcnutrition.com). In addition to its role as a precursor for neurotransmitters, tyrosine plays an important role for the function of many proteins. Within many proteins or enzymes, certain tyrosine residues can be tagged (at the hydroxyl group) with a phosphate group (phosphorylated) by specialized protein kinases. In its phosphorylated form, tyrosine is called phosphotyrosine. Tyrosine phosphorylation is considered to be one of the key steps in signal transduction and regulation of enzymatic activity. Tyrosine (or its precursor phenylalanine) is also needed to synthesize the benzoquinone structure which forms part of coenzyme Q10. |
|---|
| Structure | N[C@@H](CC1=CC=C(O)C=C1)C(O)=O InChI=1S/C9H11NO3/c10-8(9(12)13)5-6-1-3-7(11)4-2-6/h1-4,8,11H,5,10H2,(H,12,13)/t8-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (-)-alpha-Amino-p-hydroxyhydrocinnamic acid | ChEBI | | (2S)-2-Amino-3-(4-hydroxyphenyl)propanoic acid | ChEBI | | (S)-(-)-Tyrosine | ChEBI | | (S)-2-Amino-3-(p-hydroxyphenyl)propionic acid | ChEBI | | (S)-3-(p-Hydroxyphenyl)alanine | ChEBI | | (S)-alpha-Amino-4-hydroxybenzenepropanoic acid | ChEBI | | (S)-Tyrosine | ChEBI | | 4-Hydroxy-L-phenylalanine | ChEBI | | L-Tyrosin | ChEBI | | Tyr | ChEBI | | TYROSINE | ChEBI | | Y | ChEBI | | (-)-a-Amino-p-hydroxyhydrocinnamate | Generator | | (-)-a-Amino-p-hydroxyhydrocinnamic acid | Generator | | (-)-alpha-Amino-p-hydroxyhydrocinnamate | Generator | | (-)-Α-amino-p-hydroxyhydrocinnamate | Generator | | (-)-Α-amino-p-hydroxyhydrocinnamic acid | Generator | | (2S)-2-Amino-3-(4-hydroxyphenyl)propanoate | Generator | | (S)-2-Amino-3-(p-hydroxyphenyl)propionate | Generator | | (S)-a-Amino-4-hydroxybenzenepropanoate | Generator | | (S)-a-Amino-4-hydroxybenzenepropanoic acid | Generator | | (S)-alpha-Amino-4-hydroxybenzenepropanoate | Generator | | (S)-Α-amino-4-hydroxybenzenepropanoate | Generator | | (S)-Α-amino-4-hydroxybenzenepropanoic acid | Generator | | (S)-a-Amino-4-hydroxy-benzenepropanoate | HMDB | | (S)-a-Amino-4-hydroxy-benzenepropanoic acid | HMDB | | (S)-alpha-Amino-4-hydroxy-benzenepropanoate | HMDB | | (S)-alpha-Amino-4-hydroxy-benzenepropanoic acid | HMDB | | 2-Amino-3-(4-hydroxyphen yl)-2-amino-3-(4-hydroxyphenyl)-propanoate | HMDB | | 2-Amino-3-(4-hydroxyphen yl)-2-amino-3-(4-hydroxyphenyl)-propanoic acid | HMDB | | 3-(4-Hydroxyphenyl)-L-alanine | HMDB | | Benzenepropanoate | HMDB | | Benzenepropanoic acid | HMDB | | L-p-Tyrosine | HMDB | | p-Tyrosine | HMDB | | L Tyrosine | HMDB | | Tyrosine, L-isomer | HMDB | | Tyrosine, L isomer | HMDB | | Para tyrosine | HMDB | | Para-tyrosine | HMDB |
|
|---|
| Chemical Formula | C9H11NO3 |
|---|
| Average Molecular Weight | 181.1885 |
|---|
| Monoisotopic Molecular Weight | 181.073893223 |
|---|
| IUPAC Name | (2S)-2-amino-3-(4-hydroxyphenyl)propanoic acid |
|---|
| Traditional Name | L-tyrosine |
|---|
| CAS Registry Number | 60-18-4 |
|---|
| SMILES | N[C@@H](CC1=CC=C(O)C=C1)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C9H11NO3/c10-8(9(12)13)5-6-1-3-7(11)4-2-6/h1-4,8,11H,5,10H2,(H,12,13)/t8-/m0/s1 |
|---|
| InChI Key | OUYCCCASQSFEME-QMMMGPOBSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as tyrosine and derivatives. Tyrosine and derivatives are compounds containing tyrosine or a derivative thereof resulting from reaction of tyrosine at the amino group or the carboxy group, or from the replacement of any hydrogen of glycine by a heteroatom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Tyrosine and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Tyrosine or derivatives
- Phenylalanine or derivatives
- 3-phenylpropanoic-acid
- Alpha-amino acid
- Amphetamine or derivatives
- L-alpha-amino acid
- 1-hydroxy-2-unsubstituted benzenoid
- Phenol
- Aralkylamine
- Monocyclic benzene moiety
- Benzenoid
- Amino acid
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Organic oxide
- Organooxygen compound
- Organonitrogen compound
- Amine
- Primary aliphatic amine
- Organic nitrogen compound
- Carbonyl group
- Organopnictogen compound
- Organic oxygen compound
- Hydrocarbon derivative
- Primary amine
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | Not Available |
|---|
| Process | Not Available |
|---|
| Role | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 343 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 0.48 mg/mL | Not Available | | LogP | -2.26 | HANSCH,C ET AL. (1995) |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 1.62 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 9.4301 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 7.78 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 334.1 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 491.3 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 276.6 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 77.1 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 165.3 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 62.5 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 269.6 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 245.9 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 786.7 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 626.4 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 102.5 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 688.0 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 161.2 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 194.1 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 582.4 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 446.8 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 310.5 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| L-Tyrosine,1TMS,isomer #1 | C[Si](C)(C)OC1=CC=C(C[C@H](N)C(=O)O)C=C1 | 1902.3 | Semi standard non polar | 33892256 | | L-Tyrosine,1TMS,isomer #2 | C[Si](C)(C)OC(=O)[C@@H](N)CC1=CC=C(O)C=C1 | 1913.3 | Semi standard non polar | 33892256 | | L-Tyrosine,1TMS,isomer #3 | C[Si](C)(C)N[C@@H](CC1=CC=C(O)C=C1)C(=O)O | 1994.1 | Semi standard non polar | 33892256 | | L-Tyrosine,2TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@@H](N)CC1=CC=C(O[Si](C)(C)C)C=C1 | 1880.7 | Semi standard non polar | 33892256 | | L-Tyrosine,2TMS,isomer #2 | C[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C)C=C1)C(=O)O | 1957.7 | Semi standard non polar | 33892256 | | L-Tyrosine,2TMS,isomer #3 | C[Si](C)(C)N[C@@H](CC1=CC=C(O)C=C1)C(=O)O[Si](C)(C)C | 1961.5 | Semi standard non polar | 33892256 | | L-Tyrosine,2TMS,isomer #4 | C[Si](C)(C)N([C@@H](CC1=CC=C(O)C=C1)C(=O)O)[Si](C)(C)C | 2154.9 | Semi standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C)C=C1)C(=O)O[Si](C)(C)C | 1931.2 | Semi standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C)C=C1)C(=O)O[Si](C)(C)C | 1957.0 | Standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C)C=C1)C(=O)O[Si](C)(C)C | 2113.8 | Standard polar | 33892256 | | L-Tyrosine,3TMS,isomer #2 | C[Si](C)(C)OC1=CC=C(C[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C)C=C1 | 2120.2 | Semi standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #2 | C[Si](C)(C)OC1=CC=C(C[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C)C=C1 | 2016.5 | Standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #2 | C[Si](C)(C)OC1=CC=C(C[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C)C=C1 | 2235.0 | Standard polar | 33892256 | | L-Tyrosine,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O)C=C1)N([Si](C)(C)C)[Si](C)(C)C | 2093.2 | Semi standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O)C=C1)N([Si](C)(C)C)[Si](C)(C)C | 2107.1 | Standard non polar | 33892256 | | L-Tyrosine,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O)C=C1)N([Si](C)(C)C)[Si](C)(C)C | 2231.2 | Standard polar | 33892256 | | L-Tyrosine,4TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O[Si](C)(C)C)C=C1)N([Si](C)(C)C)[Si](C)(C)C | 2149.4 | Semi standard non polar | 33892256 | | L-Tyrosine,4TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O[Si](C)(C)C)C=C1)N([Si](C)(C)C)[Si](C)(C)C | 2067.2 | Standard non polar | 33892256 | | L-Tyrosine,4TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O[Si](C)(C)C)C=C1)N([Si](C)(C)C)[Si](C)(C)C | 2062.0 | Standard polar | 33892256 | | L-Tyrosine,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC1=CC=C(C[C@H](N)C(=O)O)C=C1 | 2161.7 | Semi standard non polar | 33892256 | | L-Tyrosine,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC1=CC=C(O)C=C1 | 2152.1 | Semi standard non polar | 33892256 | | L-Tyrosine,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CC=C(O)C=C1)C(=O)O | 2221.2 | Semi standard non polar | 33892256 | | L-Tyrosine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1 | 2398.0 | Semi standard non polar | 33892256 | | L-Tyrosine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)C(=O)O | 2489.8 | Semi standard non polar | 33892256 | | L-Tyrosine,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CC=C(O)C=C1)C(=O)O[Si](C)(C)C(C)(C)C | 2399.4 | Semi standard non polar | 33892256 | | L-Tyrosine,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CC=C(O)C=C1)C(=O)O)[Si](C)(C)C(C)(C)C | 2554.8 | Semi standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)C(=O)O[Si](C)(C)C(C)(C)C | 2658.8 | Semi standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)C(=O)O[Si](C)(C)C(C)(C)C | 2577.0 | Standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)C(=O)O[Si](C)(C)C(C)(C)C | 2488.8 | Standard polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC1=CC=C(C[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C=C1 | 2858.8 | Semi standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC1=CC=C(C[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C=C1 | 2627.1 | Standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC1=CC=C(C[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C)C=C1 | 2538.5 | Standard polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O)C=C1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2747.5 | Semi standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O)C=C1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2706.4 | Standard non polar | 33892256 | | L-Tyrosine,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O)C=C1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2523.4 | Standard polar | 33892256 | | L-Tyrosine,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 3059.3 | Semi standard non polar | 33892256 | | L-Tyrosine,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2824.6 | Standard non polar | 33892256 | | L-Tyrosine,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CC=C(O[Si](C)(C)C(C)(C)C)C=C1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2499.2 | Standard polar | 33892256 |
|
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-014i-0690000000-cbbf40bb26fc84f2aead | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-014i-0890000000-ca45f993f95c8b0cee44 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-014i-0890000000-5749069211ba15d713ef | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-00xr-9240000000-2c87373c0d964e0edef5 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-014i-0890000000-848b2a4f247a0b3f14e8 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-00xr-9450000000-6d4550940f4dde6f18ff | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-MS (2 TMS) | splash10-004i-1910000000-5cc19cad5dc24b3b9b11 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-MS (3 TMS) | splash10-014i-1790000000-de22041357aadf60a06b | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Non-derivatized) | splash10-014i-0690000000-cbbf40bb26fc84f2aead | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Non-derivatized) | splash10-014i-0890000000-ca45f993f95c8b0cee44 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Non-derivatized) | splash10-014i-0890000000-5749069211ba15d713ef | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Non-derivatized) | splash10-00xr-9240000000-2c87373c0d964e0edef5 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Non-derivatized) | splash10-014i-0890000000-848b2a4f247a0b3f14e8 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-QQ (Non-derivatized) | splash10-0udi-3319000000-1d3d28a67f82366fff22 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-EI-TOF (Non-derivatized) | splash10-00xr-9450000000-6d4550940f4dde6f18ff | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-MS (Non-derivatized) | splash10-004i-1910000000-5cc19cad5dc24b3b9b11 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Tyrosine GC-MS (Non-derivatized) | splash10-014i-1790000000-de22041357aadf60a06b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (Non-derivatized) - 70eV, Positive | splash10-052r-4900000000-9be1412408207db5df4e | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (2 TMS) - 70eV, Positive | splash10-05fr-9750000000-937b6ee7a745865ee7ee | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (TMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Tyrosine GC-MS (TMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0a4i-3900000000-7a26097fda66f2f445b5 | 2018-05-25 | Not Available | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-01p9-0900000000-580de2c16cd24559cd5c | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-0006-9400000000-f233fbc6c58236ee4aeb | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-002f-9100000000-8b5e12eba034bcfdf8a1 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0920000000-b43356ad3da227b488cb | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-014i-0900000000-c04f0be6515621dda5ac | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-0900000000-3ed8b68bfade9194763b | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0udi-0900000000-fe6a1ce69851a8c5db00 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0900000000-aafdcea07be221817fd4 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0002-0900000000-66a5b9a0a48bdc0a2b47 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-014i-0900000000-13eb4252ca23455a58da | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-014i-0900000000-fb85798746829bac3f3d | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03e9-0839226000-5504e667281c746669ec | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03di-0900000000-f65cb3ad2fa730c922f7 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0900000000-a9276fe43ef61b4693e6 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0a59-0039210000-21b4bd9870bf4965a6d1 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0848491200-aae99eb0b66dd1c7036d | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03di-0900000000-4104a2ca5d5ef5f22f6d | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0900000000-f309996d57a95c719deb | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03di-0013090000-112cd9c2eea42dbc079b | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-001i-0900000000-c7f95918d936586f633d | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-03yi-1900000000-d1682546c1e0893c71e4 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-014i-2900000000-a0cc78ed35e56dd812a5 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-00kf-9500000000-d3f399f5dd10e338e25a | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative-QTOF | splash10-0006-9200000000-8acd8d370f194bfe28ed | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Tyrosine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive-QTOF | splash10-001i-0900000000-6b26ce2f5b326ace12a4 | 2012-08-31 | HMDB team, MONA | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 400 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Mitochondria
|
|---|
| Biospecimen Locations | - Blood
- Breast Milk
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Sweat
- Urine
|
|---|
| Tissue Locations | - All Tissues
- Placenta
- Prostate
|
|---|
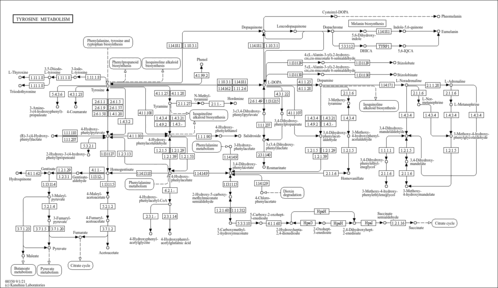
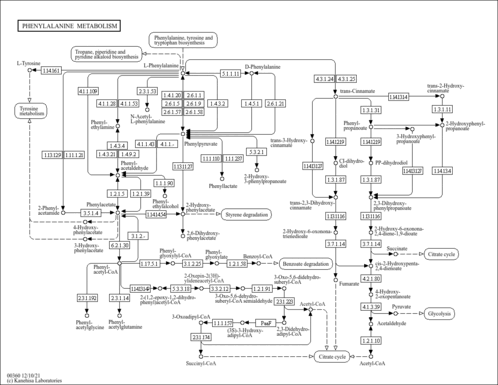
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 59.0 (47.0-71.0) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 120.0 +/- 100.0 uM | Newborn (0-30 days old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 68.0 +/- 10.0 uM | Children (1-13 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 72.0 +/- 15.0 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 61.0 +/- 13.0 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 142.0 +/- 4.0 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 54.5 +/- 9.7 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 143 +/- 35.3 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 32-124 uM | Newborn (0-30 days old) | Both | Normal | | details | | Blood | Detected and Quantified | 29-135 uM | Infant (1 - 3 months old) | Both | Normal | | details | | Blood | Detected and Quantified | 24-105 uM | Children (3 months - 6 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 43-88 uM | Children (6 - 18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 65.1 +/- 2.55 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 62.9 +/- 12.2 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 35.0-80.0 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 55 - 147 uM | Newborn (0-30 days old) | Both | Normal | | details | | Blood | Detected and Quantified | 55-147 uM | Infant (0-1 year old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 46 +/- 8 uM | Children (1 - 13 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 61 +/- 13 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 72 +/- 15 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 60.8(48.3-71.9) uM | Children (1-13 uears old) | Both | Normal | | details | | Blood | Detected and Quantified | 84.2 (67.5-109) uM | Infant (0-1 year old) | Not Available | Normal | | details | | Blood | Detected and Quantified | 81.9 (69.8-94) uM | Newborn (0-30 days old) | Not Available | Normal | | details | | Blood | Detected and Quantified | 34-94 uM | Newborn (0-30 days old) | Not Specified | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 66.49 ± 7.87 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 93.04 ± 21.72 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 85.11 ± 30.2 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 73.31 ± 20.35 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 82.41 ± 28.24 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 77.13 ± 27.07 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 72.67 ± 16.68 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 88.24 ± 23.04 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 70.02 ± 26.53 uM | Adult (>18 years old) | Both | Normal | | details | | Breast Milk | Detected and Quantified | 15.0 +/- 6.6 uM | Adult (>18 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 6.4 +/- 1.5 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 10.1 (6.37-13.8) uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 24.5 +/- 8.9 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 9.3 +/- 2.0 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 7.9 +/- 3.0 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 12.0 +/- 9.0 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 8.0 +/- 1.0 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 12.2 +/- 3.8 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 3.9-18.5 uM | Children (0 - 10 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 3.9-17 uM | Adolescent (>11 years old) | Both | Normal | | details | | Feces | Detected and Quantified | 114.246 +/- 94.322 nmol/g wet feces | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Both | Normal | | details | | Feces | Detected and Quantified | 478.323 (294.353-662.294) nmol/g wet feces | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected and Quantified | 1048.632 (588.706-1508.558) nmol/g wet feces | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected and Quantified | 1453.367 (607.103-2299.631) nmol/g wet feces | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected and Quantified | 700 +/- 420 nmol/g wet feces | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected and Quantified | 110 +/- 100 nmol/g wet feces | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | >10 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 1-127 uM | Adult (>18 years old) | Male | normal | | details | | Saliva | Detected and Quantified | 1-68 uM | Adult (>18 years old) | Male | normal | | details | | Saliva | Detected and Quantified | 39.85 +/- 24.71 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 13.7 +/- 3.6 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 12.0 +/- 5.8 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 11.8 +/- 6.2 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 12.2 +/- 3.8 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 5 (<1-78) uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 1-77 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 30 (<1-91) uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 12 (<1-49) uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 36.17 +/- 15.02 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 11.1 +/- 7.65 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 15.1 +/- 10.3 uM | Adult (>18 years old) | Female | Normal | | details | | Sweat | Detected and Quantified | 70-270 uM | Adult (60 years old) | Male | Normal | | details | | Sweat | Detected and Quantified | 40 uM | Adult (40 years old) | Male | Normal | | details | | Sweat | Detected but not Quantified | Not Quantified | Adult | Both | Normal | | details | | Urine | Detected and Quantified | 27.35 +/- 13.98 umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 10.9 (2.566-19.1) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.72 (0.18-1.38) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 4.2 +/- 1.85 umol/mmol creatinine | Children (1-13 years old) | Male | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 8.8 +/- 4.5 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 7.0 +/- 3.0 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 21.0 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 6.72 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 5.52-38.66 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 7.41 +/- 2.30 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 7.5 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 8.7 (3.0-20.1) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 15.1 (6.1-23.2) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 9.5 (4.1-23.5) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 24.3893 +/- 10.0806 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 14.588 +/- 10.744 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 6.22 - 67.72 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 13.79 - 63.88 umol/mmol creatinine | Infant (1 - 6 months old) | Both | Normal | | details | | Urine | Detected and Quantified | 12.55 - 66.48 umol/mmol creatinine | Infant (6 months - <1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 15.04 - 59.02 umol/mmol creatinine | Children (1 - 2 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 9.95 - 29.96 umol/mmol creatinine | Children (2 - 4 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 6.22 - 42.62 umol/mmol creatinine | Children (4 - 13 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.36 - 27.70 umol/mmol creatinine | Adolescent (13 - 21 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.36 - 27.70 umol/mmol creatinine | Adult (>21 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 4.3-9.2 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 6.1-11.1 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 15.0370-168.459 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.0 +/- 0.5 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.9 +/- 0.7 umol/mmol creatinine | Infant (1 - 6 months old) | Both | Normal | | details | | Urine | Detected and Quantified | 2.7 +/- 0.8 umol/mmol creatinine | Infant (6 months - <1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 42.96 +/- 27.13 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 66.71 +/- 47.49 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 70.10 +/- 45.22 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 6-57 umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 2.1-14.81 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 5.82 +/- 3.26 umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected and Quantified | 5.96 +/- 2.98 umol/mmol creatinine | Newborn (0-30 days old) | Male | Normal | | details | | Urine | Detected and Quantified | 4.42-77.47 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 4.30-54.18 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 2.60-28.73 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 2.49-27.71 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 1.36-23.53 umol/mmol creatinine | Adolescent (13-18 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 1.70-13.0086 umol/mmol creatinine | Adult (>18 years old) | Not Specified | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 5.03 (2.60-7.72) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details |
|
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 45.63 (19.15) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Oesophageal cancer | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected and Quantified | 57.2 +/- 3.33 uM | Adult (>18 years old) | Both | Schizophrenia | | details | | Blood | Detected and Quantified | 54.1 (15.3) uM | Adult (>18 years old) | Female | Early preeclampsia | | details | | Blood | Detected and Quantified | 58.0 (22.3) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 65.1 (23.7) uM | Adult (>18 years old) | Female | Late-onset preeclampsia | | details | | Blood | Detected and Quantified | 62.3 (21.7) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 50.5 (15.4) uM | Adult (>18 years old) | Female | Down syndrome pregnancy | | details | | Blood | Detected and Quantified | 54.1 (20.7) uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 56.9 +/- 15.2 uM | Adult (>18 years old) | Female | Pregnancy with fetuses with trisomy 18 | | details | | Blood | Detected and Quantified | 55.82 +/- 19.6 uM | Adult (>18 years old) | Female | Pregnancy | | details | | Blood | Detected and Quantified | 42.94 (11.38) uM | Adult (>18 years old) | Female | Pregnancy with fetus having congenital heart defect | | details | | Blood | Detected and Quantified | 50.7 (46.8-54.6) uM | Adult (>18 years old) | Both | Epilepsy | | details | | Blood | Detected and Quantified | 257 uM | Infant (0-1 year old) | Male | Fumaric aciduria | | details | | Blood | Detected and Quantified | 1471.948 uM | Children (1 - 13 years old) | Male | Hereditary tyrosinemia | | details | | Blood | Detected and Quantified | >220 uM | Newborn (0-30 days old) | Both | Tyrosinemia I | | details | | Blood | Detected and Quantified | 108-146 uM | Children (1-13 years old) | Female | S-Adenosylhomocysteine hydrolase deficiency | | details | | Blood | Detected and Quantified | 310-1410 uM | Children (1-13 years old) | Both | Tyrosinemia I | | details | | Blood | Detected and Quantified | 395.1 - 586.7 uM | Newborn (0-30 days old) | Not Available | Hawkinsinuria | | details | | Blood | Detected and Quantified | 48-95 uM | Children (1-13 years old) | Female | S-Adenosylhomocysteine hydrolase deficiency | | details | | Blood | Detected and Quantified | 61.1(52.0-72.9) uM | Children (1-13 uears old) | Both | Environmental enteric dysfunction | | details | | Blood | Detected and Quantified | 82.5 +/- 21.1 uM | Children (1-13 years old) | Both | Obesity | | details | | Blood | Detected and Quantified | 91.7 +/- 19.9 uM | Children (1-13 years old) | Both | Obesity | | details | | Blood | Detected and Quantified | 143-246 uM | Newborn (0-30 days old) | Both | Citrullinemia type II, neonatal-onset | | details | | Blood | Detected and Quantified | 108(80-350) uM | Adult (>18 years old) | Both | Sepsis | | details | | Blood | Detected and Quantified | 74.0 (71.0-77.0) uM | Adult (>18 years old) | Both | Myocardial infarction | | details | | Blood | Detected and Quantified | 81.0 (75.9-86.1) uM | Adult (>18 years old) | Both | Epilepsy | | details | | Blood | Detected and Quantified | 90.0 (81.0-99.0) uM | Adult (>18 years old) | Both | Viral Infection | | details | | Blood | Detected and Quantified | 57.2 +/- 24.4 uM | Adult (>18 years old) | Both | Heart Transplant | | details | | Blood | Detected and Quantified | 143.77 +/- 15.49 uM | Elderly (>65 years old) | Both | Alzheimer's disease | | details | | Blood | Detected and Quantified | 73.0 +/- 20.0 uM | Adult (>18 years old) | Female | Schizophrenia | | details | | Blood | Detected and Quantified | 91.2 +/- 30.0 uM | Adult (>18 years old) | Male | Schizophrenia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 9.4 +/- 4.9 uM | Adult (>18 years old) | Not Specified | Epilepsy | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 27.18 +/- 4.8 uM | Adult (>18 years old) | Both | Alzheimer's disease | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 7.8 +/- 1.38 uM | Adult (>18 years old) | Both | Schizophrenia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 15.3 +/- 4.9 uM | Adult (>18 years old) | Both | Hypothyroidism | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 15.3 +/- 4.9 uM | Adult (>18 years old) | Not Specified | hypothyroidism | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 12.6 +/- 3.3 uM | Children (1-13 years old) | Both | Leukemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 13.0 +/- 4.1 uM | Children (1-13 years old) | Both | Leukemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 8.05 +/- 3.03 uM | Adult (>18 years old) | Not Specified | Epilepsy | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 12.3 +/- 9.3 uM | Adult (>18 years old) | Not Specified | Epilepsy | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 13.0 +/- 10.6 uM | Adult (>18 years old) | Not Specified | Epilepsy | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 14.2 +/-12.5 uM | Adult (>18 years old) | Not Specified | Epilepsy | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 15.3 (10.4-20.2) uM | Adult (>18 years old) | Both | Hypothyroidism | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Unclassified IBD | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Myalgic encephalomyelitis/chronic fatigue syndrome | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | CCD | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Ileal Crohn's disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Irritable Bowel Syndrome | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Autism | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Irritable bowel syndrome | | details | | Saliva | Detected and Quantified | 25.82 +/- 9.25 uM | Adult (>18 years old) | Male | Frontotemporal lobe dementia | | details | | Saliva | Detected and Quantified | 25.05 +/- 8.02 uM | Adult (>18 years old) | Both | Lewy body disease | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Periodontal Probing Depth | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Missing teeth | | details | | Saliva | Detected and Quantified | 21.64 +/- 9.24 uM | Adult (>18 years old) | Male | Alzheimer's disease | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Periodontal diseases | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Pancreatic cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Attachment loss | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Breast cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Oral cancer | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Bladder cancer | | details | | Urine | Detected and Quantified | 10.72 +/- 4.63 umol/mmol creatinine | Adult (>18 years old) | Not Specified | Cachexia | | details | | Urine | Detected and Quantified | 35.150-369 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Fumaric aciduria | | details | | Urine | Detected and Quantified | 10.5 +/- 1.5 umol/mmol creatinine | Adult (>18 years old) | Both | Alzheimer's disease | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Urine | Detected and Quantified | 16.005 +/- 10.562 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected and Quantified | 16.2462 +/- 14.3236 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Gastroesophageal reflux disease | | details | | Urine | Detected and Quantified | 20.2606 +/- 11.9167 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details |
|
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Colorectal cancer |
|---|
- Qiu Y, Cai G, Su M, Chen T, Zheng X, Xu Y, Ni Y, Zhao A, Xu LX, Cai S, Jia W: Serum metabolite profiling of human colorectal cancer using GC-TOFMS and UPLC-QTOFMS. J Proteome Res. 2009 Oct;8(10):4844-50. doi: 10.1021/pr9004162. [PubMed:19678709 ]
- Ni Y, Xie G, Jia W: Metabonomics of human colorectal cancer: new approaches for early diagnosis and biomarker discovery. J Proteome Res. 2014 Sep 5;13(9):3857-70. doi: 10.1021/pr500443c. Epub 2014 Aug 14. [PubMed:25105552 ]
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
| | Sepsis |
|---|
- Ferrario M, Cambiaghi A, Brunelli L, Giordano S, Caironi P, Guatteri L, Raimondi F, Gattinoni L, Latini R, Masson S, Ristagno G, Pastorelli R: Mortality prediction in patients with severe septic shock: a pilot study using a target metabolomics approach. Sci Rep. 2016 Feb 5;6:20391. doi: 10.1038/srep20391. [PubMed:26847922 ]
| | Citrullinemia type II, neonatal-onset |
|---|
- Ohura T, Kobayashi K, Tazawa Y, Nishi I, Abukawa D, Sakamoto O, Iinuma K, Saheki T: Neonatal presentation of adult-onset type II citrullinemia. Hum Genet. 2001 Feb;108(2):87-90. [PubMed:11281457 ]
| | Obesity |
|---|
- Simone Wahl, Christina Holzapfel, Zhonghao Yu, Michaela Breier, Ivan Kondofersky, Christiane Fuchs, Paula Singmann, Cornelia Prehn, Jerzy Adamski, Harald Grallert, Thomas Illig, Rui Wang-Sattler, Thomas Reinehr (2013). Metabolomics reveals determinants of weight loss during lifestyle intervention in obese children. Metabolomics.
| | Hawkinsinuria |
|---|
- Thodi G, Schulpis KH, Dotsikas Y, Pavlides C, Molou E, Chatzidaki M, Triantafylli O, Loukas YL: Hawkinsinuria in two unrelated Greek newborns: identification of a novel variant, biochemical findings and treatment. J Pediatr Endocrinol Metab. 2016 Jan;29(1):15-20. doi: 10.1515/jpem-2015-0132. [PubMed:26226126 ]
| | Hypermethioninemia |
|---|
- Labrune P, Perignon JL, Rault M, Brunet C, Lutun H, Charpentier C, Saudubray JM, Odievre M: Familial hypermethioninemia partially responsive to dietary restriction. J Pediatr. 1990 Aug;117(2 Pt 1):220-6. [PubMed:2380820 ]
| | Tyrosinemia I |
|---|
- Nakamura K, Matsumoto S, Mitsubuchi H, Endo F: Diagnosis and treatment of hereditary tyrosinemia in Japan. Pediatr Int. 2015;57(1):37-40. doi: 10.1111/ped.12550. [PubMed:25443793 ]
- Gissen P, Preece MA, Willshaw HA, McKiernan PJ: Ophthalmic follow-up of patients with tyrosinaemia type I on NTBC. J Inherit Metab Dis. 2003;26(1):13-6. [PubMed:12872835 ]
| | Tyrosinemia |
|---|
- Swarna M, Jyothy A, Usha Rani P, Reddy PP: Amino acid disorders in mental retardation: a two-decade study from Andhra Pradesh. Biochem Genet. 2004 Apr;42(3-4):85-98. [PubMed:15168722 ]
| | Fumarase deficiency |
|---|
- Allegri G, Fernandes MJ, Scalco FB, Correia P, Simoni RE, Llerena JC Jr, de Oliveira ML: Fumaric aciduria: an overview and the first Brazilian case report. J Inherit Metab Dis. 2010 Aug;33(4):411-9. doi: 10.1007/s10545-010-9134-2. Epub 2010 Jun 15. [PubMed:20549362 ]
| | Late-onset preeclampsia |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: First-trimester metabolomic detection of late-onset preeclampsia. Am J Obstet Gynecol. 2013 Jan;208(1):58.e1-7. doi: 10.1016/j.ajog.2012.11.003. Epub 2012 Nov 13. [PubMed:23159745 ]
| | Pregnancy |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomics and first-trimester prediction of early-onset preeclampsia. J Matern Fetal Neonatal Med. 2012 Oct;25(10):1840-7. doi: 10.3109/14767058.2012.680254. Epub 2012 Apr 28. [PubMed:22494326 ]
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: First-trimester metabolomic detection of late-onset preeclampsia. Am J Obstet Gynecol. 2013 Jan;208(1):58.e1-7. doi: 10.1016/j.ajog.2012.11.003. Epub 2012 Nov 13. [PubMed:23159745 ]
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomic analysis for first-trimester Down syndrome prediction. Am J Obstet Gynecol. 2013 May;208(5):371.e1-8. doi: 10.1016/j.ajog.2012.12.035. Epub 2013 Jan 8. [PubMed:23313728 ]
- Bahado-Singh RO, Akolekar R, Chelliah A, Mandal R, Dong E, Kruger M, Wishart DS, Nicolaides K: Metabolomic analysis for first-trimester trisomy 18 detection. Am J Obstet Gynecol. 2013 Jul;209(1):65.e1-9. doi: 10.1016/j.ajog.2013.03.028. Epub 2013 Mar 25. [PubMed:23535240 ]
- Bahado-Singh RO, Ertl R, Mandal R, Bjorndahl TC, Syngelaki A, Han B, Dong E, Liu PB, Alpay-Savasan Z, Wishart DS, Nicolaides KH: Metabolomic prediction of fetal congenital heart defect in the first trimester. Am J Obstet Gynecol. 2014 Sep;211(3):240.e1-240.e14. doi: 10.1016/j.ajog.2014.03.056. Epub 2014 Apr 1. [PubMed:24704061 ]
| | Early preeclampsia |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomics and first-trimester prediction of early-onset preeclampsia. J Matern Fetal Neonatal Med. 2012 Oct;25(10):1840-7. doi: 10.3109/14767058.2012.680254. Epub 2012 Apr 28. [PubMed:22494326 ]
| | Epilepsy |
|---|
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
- Botez MI, Young SN: Effects of anticonvulsant treatment and low levels of folate and thiamine on amine metabolites in cerebrospinal fluid. Brain. 1991 Feb;114 ( Pt 1A):333-48. [PubMed:1705463 ]
| | Myocardial infarction |
|---|
- Wannemacher RW Jr, Klainer AS, Dinterman RE, Beisel WR: The significance and mechanism of an increased serum phenylalanine-tyrosine ratio during infection. Am J Clin Nutr. 1976 Sep;29(9):997-1006. [PubMed:822705 ]
| | Viral infection |
|---|
- Wannemacher RW Jr, Klainer AS, Dinterman RE, Beisel WR: The significance and mechanism of an increased serum phenylalanine-tyrosine ratio during infection. Am J Clin Nutr. 1976 Sep;29(9):997-1006. [PubMed:822705 ]
| | Schizophrenia |
|---|
- Alfredsson G, Wiesel FA: Monoamine metabolites and amino acids in serum from schizophrenic patients before and during sulpiride treatment. Psychopharmacology (Berl). 1989;99(3):322-7. [PubMed:2480613 ]
- Do KQ, Lauer CJ, Schreiber W, Zollinger M, Gutteck-Amsler U, Cuenod M, Holsboer F: gamma-Glutamylglutamine and taurine concentrations are decreased in the cerebrospinal fluid of drug-naive patients with schizophrenic disorders. J Neurochem. 1995 Dec;65(6):2652-62. [PubMed:7595563 ]
- Fukushima T, Iizuka H, Yokota A, Suzuki T, Ohno C, Kono Y, Nishikiori M, Seki A, Ichiba H, Watanabe Y, Hongo S, Utsunomiya M, Nakatani M, Sadamoto K, Yoshio T: Quantitative analyses of schizophrenia-associated metabolites in serum: serum D-lactate levels are negatively correlated with gamma-glutamylcysteine in medicated schizophrenia patients. PLoS One. 2014 Jul 8;9(7):e101652. doi: 10.1371/journal.pone.0101652. eCollection 2014. [PubMed:25004141 ]
| | Alzheimer's disease |
|---|
- Fonteh AN, Harrington RJ, Tsai A, Liao P, Harrington MG: Free amino acid and dipeptide changes in the body fluids from Alzheimer's disease subjects. Amino Acids. 2007 Feb;32(2):213-24. Epub 2006 Oct 10. [PubMed:17031479 ]
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Hypothyroidism |
|---|
- Sjoberg S, Eriksson M, Nordin C: L-thyroxine treatment and neurotransmitter levels in the cerebrospinal fluid of hypothyroid patients: a pilot study. Eur J Endocrinol. 1998 Nov;139(5):493-7. [PubMed:9849813 ]
| | Leukemia |
|---|
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [PubMed:15911239 ]
| | Irritable bowel syndrome |
|---|
- Le Gall G, Noor SO, Ridgway K, Scovell L, Jamieson C, Johnson IT, Colquhoun IJ, Kemsley EK, Narbad A: Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011 Sep 2;10(9):4208-18. doi: 10.1021/pr2003598. Epub 2011 Aug 8. [PubMed:21761941 ]
- Hong YS, Hong KS, Park MH, Ahn YT, Lee JH, Huh CS, Lee J, Kim IK, Hwang GS, Kim JS: Metabonomic understanding of probiotic effects in humans with irritable bowel syndrome. J Clin Gastroenterol. 2011 May-Jun;45(5):415-25. doi: 10.1097/MCG.0b013e318207f76c. [PubMed:21494186 ]
| | Crohn's disease |
|---|
- Bjerrum JT, Wang Y, Hao F, Coskun M, Ludwig C, Gunther U, Nielsen OH: Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn's disease and healthy individuals. Metabolomics. 2015;11:122-133. Epub 2014 Jun 1. [PubMed:25598765 ]
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Autism |
|---|
- De Angelis M, Piccolo M, Vannini L, Siragusa S, De Giacomo A, Serrazzanetti DI, Cristofori F, Guerzoni ME, Gobbetti M, Francavilla R: Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS One. 2013 Oct 9;8(10):e76993. doi: 10.1371/journal.pone.0076993. eCollection 2013. [PubMed:24130822 ]
| | Ulcerative colitis |
|---|
- Le Gall G, Noor SO, Ridgway K, Scovell L, Jamieson C, Johnson IT, Colquhoun IJ, Kemsley EK, Narbad A: Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011 Sep 2;10(9):4208-18. doi: 10.1021/pr2003598. Epub 2011 Aug 8. [PubMed:21761941 ]
- Bjerrum JT, Wang Y, Hao F, Coskun M, Ludwig C, Gunther U, Nielsen OH: Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn's disease and healthy individuals. Metabolomics. 2015;11:122-133. Epub 2014 Jun 1. [PubMed:25598765 ]
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Periodontal disease |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Pancreatic cancer |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Perillyl alcohol administration for cancer treatment |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Frontotemporal dementia |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Attachment loss |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Missing teeth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Periodontal Probing Depth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Lewy body disease |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Cachexia |
|---|
- Wishart DS, Knox C, Guo AC, Eisner R, Young N, Gautam B, Hau DD, Psychogios N, Dong E, Bouatra S, Mandal R, Sinelnikov I, Xia J, Jia L, Cruz JA, Lim E, Sobsey CA, Shrivastava S, Huang P, Liu P, Fang L, Peng J, Fradette R, Cheng D, Tzur D, Clements M, Lewis A, De Souza A, Zuniga A, Dawe M, Xiong Y, Clive D, Greiner R, Nazyrova A, Shaykhutdinov R, Li L, Vogel HJ, Forsythe I: HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res. 2009 Jan;37(Database issue):D603-10. doi: 10.1093/nar/gkn810. Epub 2008 Oct 25. [PubMed:18953024 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | DB00135 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB000446 |
|---|
| KNApSAcK ID | C00001397 |
|---|
| Chemspider ID | 5833 |
|---|
| KEGG Compound ID | C00082 |
|---|
| BioCyc ID | TYR |
|---|
| BiGG ID | 33785 |
|---|
| Wikipedia Link | Tyrosine |
|---|
| METLIN ID | 34 |
|---|
| PubChem Compound | 6057 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 17895 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | TYR_L |
|---|
| MarkerDB ID | MDB00000076 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Enei, Hitoshi; Matsui, Hiroshi; Yamashita, Koichi; Okumura, Shinji; Yamada, Hideaki. Microbiological synthesis of L-tyrosine and 3,4-dihydroxyphenyl-L-alanine. I. Distribution of tyrosine phenol lyase in microorganisms. Agricultural and Biological Chemist 1972, 36:11, 1861-1868, DOI: 10.1080/00021369.1972.10860505 |
|---|
| Material Safety Data Sheet (MSDS) | Download (PDF) |
|---|
| General References | - Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Silwood CJ, Lynch E, Claxson AW, Grootveld MC: 1H and (13)C NMR spectroscopic analysis of human saliva. J Dent Res. 2002 Jun;81(6):422-7. [PubMed:12097436 ]
- Nicholson JK, O'Flynn MP, Sadler PJ, Macleod AF, Juul SM, Sonksen PH: Proton-nuclear-magnetic-resonance studies of serum, plasma and urine from fasting normal and diabetic subjects. Biochem J. 1984 Jan 15;217(2):365-75. [PubMed:6696735 ]
- Engelborghs S, Marescau B, De Deyn PP: Amino acids and biogenic amines in cerebrospinal fluid of patients with Parkinson's disease. Neurochem Res. 2003 Aug;28(8):1145-50. [PubMed:12834252 ]
- Sjoberg S, Eriksson M, Nordin C: L-thyroxine treatment and neurotransmitter levels in the cerebrospinal fluid of hypothyroid patients: a pilot study. Eur J Endocrinol. 1998 Nov;139(5):493-7. [PubMed:9849813 ]
- Eklundh T, Eriksson M, Sjoberg S, Nordin C: Monoamine precursors, transmitters and metabolites in cerebrospinal fluid: a prospective study in healthy male subjects. J Psychiatr Res. 1996 May-Jun;30(3):201-8. [PubMed:8884658 ]
- Hagenfeldt L, Bjerkenstedt L, Edman G, Sedvall G, Wiesel FA: Amino acids in plasma and CSF and monoamine metabolites in CSF: interrelationship in healthy subjects. J Neurochem. 1984 Mar;42(3):833-7. [PubMed:6198473 ]
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [PubMed:15911239 ]
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [PubMed:12297216 ]
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
- Deng C, Shang C, Hu Y, Zhang X: Rapid diagnosis of phenylketonuria and other aminoacidemias by quantitative analysis of amino acids in neonatal blood spots by gas chromatography-mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci. 2002 Jul 25;775(1):115-20. [PubMed:12101068 ]
- Flamen P, Bernheim N, Deron P, Caveliers V, Chavatte K, Franken PR, Bossuyt A: Iodine-123 alpha-methyl-l-tyrosine single-photon emission tomography for the visualization of head and neck squamous cell carcinomas. Eur J Nucl Med. 1998 Feb;25(2):177-81. [PubMed:9473267 ]
- Ishiwata K, Tsukada H, Kubota K, Nariai T, Harada N, Kawamura K, Kimura Y, Oda K, Iwata R, Ishii K: Preclinical and clinical evaluation of O-[11C]methyl-L-tyrosine for tumor imaging by positron emission tomography. Nucl Med Biol. 2005 Apr;32(3):253-62. [PubMed:15820760 ]
- Wannemacher RW Jr, Klainer AS, Dinterman RE, Beisel WR: The significance and mechanism of an increased serum phenylalanine-tyrosine ratio during infection. Am J Clin Nutr. 1976 Sep;29(9):997-1006. [PubMed:822705 ]
- Molnar GA, Wagner Z, Marko L, Ko Szegi T, Mohas M, Kocsis B, Matus Z, Wagner L, Tamasko M, Mazak I, Laczy B, Nagy J, Wittmann I: Urinary ortho-tyrosine excretion in diabetes mellitus and renal failure: evidence for hydroxyl radical production. Kidney Int. 2005 Nov;68(5):2281-7. [PubMed:16221230 ]
- Molnar GA, Nemes V, Biro Z, Ludany A, Wagner Z, Wittmann I: Accumulation of the hydroxyl free radical markers meta-, ortho-tyrosine and DOPA in cataractous lenses is accompanied by a lower protein and phenylalanine content of the water-soluble phase. Free Radic Res. 2005 Dec;39(12):1359-66. [PubMed:16298866 ]
- Hoffhines AJ, Damoc E, Bridges KG, Leary JA, Moore KL: Detection and purification of tyrosine-sulfated proteins using a novel anti-sulfotyrosine monoclonal antibody. J Biol Chem. 2006 Dec 8;281(49):37877-87. Epub 2006 Oct 17. [PubMed:17046811 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
- Freund HR, Ryan JA Jr, Fischer JE: Amino acid derangements in patients with sepsis: treatment with branched chain amino acid rich infusions. Ann Surg. 1978 Sep;188(3):423-30. doi: 10.1097/00000658-197809000-00017. [PubMed:99098 ]
- Wernly B, Lichtenauer M, Hoppe UC, Jung C: Hyperglycemia in septic patients: an essential stress survival response in all, a robust marker for risk stratification in some, to be messed with in none. J Thorac Dis. 2016 Jul;8(7):E621-4. doi: 10.21037/jtd.2016.05.24. [PubMed:27501420 ]
- Knuplez E, Marsche G: An Updated Review of Pro- and Anti-Inflammatory Properties of Plasma Lysophosphatidylcholines in the Vascular System. Int J Mol Sci. 2020 Jun 24;21(12). pii: ijms21124501. doi: 10.3390/ijms21124501. [PubMed:32599910 ]
|
|---|