| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2006-08-13 15:05:29 UTC |
|---|
| Update Date | 2022-03-07 02:49:20 UTC |
|---|
| HMDB ID | HMDB0004243 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | 12(S)-HPETE |
|---|
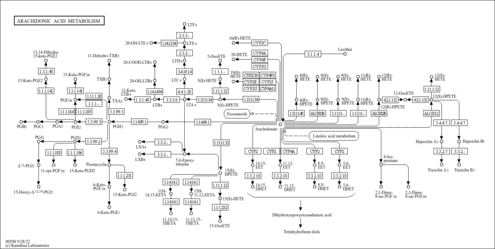
| Description | 12-HPETE is one of the six monohydroperoxy fatty acids produced by the non-enzymatic oxidation of arachidonic acid (Leukotrienes). Reduction of the hydroperoxide yields the more stable hydroxyl fatty acid (+/-)12-HETE. A family of biologically active compounds derived from arachidonic acid by oxidative metabolism through the 5-lipoxygenase pathway. They participate in host defense reactions and pathophysiological conditions such as immediate hypersensitivity and inflammation. They have potent actions on many essential organs and systems, including the cardiovascular, pulmonary, and central nervous system as well as the gastrointestinal tract and the immune system. |
|---|
| Structure | CCCCC\C=C/C[C@H](OO)\C=C\C=C/C\C=C/CCCC(O)=O InChI=1S/C20H32O4/c1-2-3-4-5-10-13-16-19(24-23)17-14-11-8-6-7-9-12-15-18-20(21)22/h7-11,13-14,17,19,23H,2-6,12,15-16,18H2,1H3,(H,21,22)/b9-7-,11-8-,13-10-,17-14+/t19-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (5Z,8Z,10E,14Z)-(12S)-12-Hydroperoxyeicosa-5,8,10,14-tetraenoate | ChEBI | | (5Z,8Z,10E,14Z)-(12S)-12-Hydroperoxyicosa-5,8,10,14-tetraenoate | ChEBI | | (5Z,8Z,10E,14Z)-(12S)-12-Hydroperoxyicosa-5,8,10,14-tetraenoic acid | ChEBI | | 12-HpETE | ChEBI | | 12-Hydroperoxyeicosatetraenoate | ChEBI | | 12-Hydroperoxyeicosatetraenoic acid | ChEBI | | 12-Hydroperoxyicosatetraenoate | ChEBI | | 12-Hydroperoxyicosatetraenoic acid | ChEBI | | 12-Oohete | ChEBI | | Arachidonic acid omega-9 hydroperoxide | ChEBI | | Omega-9 hpaa | ChEBI | | Omega-9-hydroperoxyarachidonic acid | ChEBI | | (5Z,8Z,10E,14Z)-(12S)-12-Hydroperoxyeicosa-5,8,10,14-tetraenoic acid | Generator | | Arachidonate omega-9 hydroperoxide | Generator | | Omega-9-hydroperoxyarachidonate | Generator | | (5Z,8Z,10E,12S,14Z)-12-Hydroperoxyicosa-5,8,10,14-tetraenoic acid | HMDB | | (5Z,8Z,10E,12S,14Z)-12-Hydroperoxyicosa-5,8,10,14-tetraenoate | HMDB | | 12-HPETE, (S-(e,e,e))-isomer | HMDB | | 12-L-Hydroperoxy-5,8,10,14-eicosatetraenoic acid | HMDB | | 12-Hydroperoxy-5,8,10,14-icosatetraenoic acid | HMDB | | 12-HPETE, (e,Z,Z,Z)-isomer | HMDB | | 12-HPETE, (S-(e,Z,Z,Z))-isomer | HMDB |
|
|---|
| Chemical Formula | C20H32O4 |
|---|
| Average Molecular Weight | 336.4657 |
|---|
| Monoisotopic Molecular Weight | 336.230059512 |
|---|
| IUPAC Name | (5Z,8Z,10E,12S,14Z)-12-hydroperoxyicosa-5,8,10,14-tetraenoic acid |
|---|
| Traditional Name | 12-HpETE |
|---|
| CAS Registry Number | 71774-10-2 |
|---|
| SMILES | CCCCC\C=C/C[C@H](OO)\C=C\C=C/C\C=C/CCCC(O)=O |
|---|
| InChI Identifier | InChI=1S/C20H32O4/c1-2-3-4-5-10-13-16-19(24-23)17-14-11-8-6-7-9-12-15-18-20(21)22/h7-11,13-14,17,19,23H,2-6,12,15-16,18H2,1H3,(H,21,22)/b9-7-,11-8-,13-10-,17-14+/t19-/m0/s1 |
|---|
| InChI Key | ZIOZYRSDNLNNNJ-LQWMCKPYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as hydroperoxyeicosatetraenoic acids. These are eicosanoic acids with an attached hydroperoxyl group and four CC double bonds. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Eicosanoids |
|---|
| Direct Parent | Hydroperoxyeicosatetraenoic acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Hydroperoxyeicosatetraenoic acid
- Long-chain fatty acid
- Hydroperoxy fatty acid
- Fatty acid
- Unsaturated fatty acid
- Allylic hydroperoxide
- Hydroperoxide
- Carboxylic acid derivative
- Alkyl hydroperoxide
- Carboxylic acid
- Peroxol
- Monocarboxylic acid or derivatives
- Carbonyl group
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxide
- Organic oxygen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | Not Available |
|---|
| Physical Properties |
|---|
| State | Liquid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter)). Predicted by Afia on May 17, 2022. | 7.11 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 24.071 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 1.09 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 35.0 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 3213.5 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 425.5 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 206.4 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 278.7 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 639.2 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 1032.4 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 543.8 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 115.4 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 2147.7 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 712.4 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 1717.8 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 749.1 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 493.1 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 349.6 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 526.3 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 8.6 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - 12(S)-HPETE GC-MS (Non-derivatized) - 70eV, Positive | splash10-0nml-8491000000-0127633b6ae0909f1672 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - 12(S)-HPETE GC-MS (1 TMS) - 70eV, Positive | splash10-05dr-9334000000-7ccf269dc3d82ad47d27 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - 12(S)-HPETE GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 10V, Positive-QTOF | splash10-014i-0119000000-663dc7dd460df95c9d6d | 2015-09-14 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 20V, Positive-QTOF | splash10-00pl-5975000000-e3fe517c22245902e6d2 | 2015-09-14 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 40V, Positive-QTOF | splash10-052f-9640000000-1c2eba4af6221c6ee488 | 2015-09-14 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 10V, Negative-QTOF | splash10-000i-0129000000-9b31acab4f79d4ab744c | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 20V, Negative-QTOF | splash10-014r-1659000000-5db483e2b6ae8fccb37e | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 40V, Negative-QTOF | splash10-0a4i-9740000000-a88fa00904c08bd3dbfe | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 10V, Positive-QTOF | splash10-0f79-0249000000-702a285b6044fb920d29 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 20V, Positive-QTOF | splash10-0f79-7977000000-3cd8fc7d5c58fa9ef000 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 40V, Positive-QTOF | splash10-066u-9420000000-b47740fabc6ca56ad6c0 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 10V, Negative-QTOF | splash10-000i-0009000000-7847d679fdeeabe38573 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 20V, Negative-QTOF | splash10-0udr-0329000000-d4296bbebfcef533338a | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(S)-HPETE 40V, Negative-QTOF | splash10-056u-3690000000-c9e5a737326ebfdd3bf0 | 2021-09-22 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, H2O, predicted) | 2022-08-20 | Wishart Lab | View Spectrum |
|
|---|