| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Expected but not Quantified |
|---|
| Creation Date | 2006-08-14 00:36:18 UTC |
|---|
| Update Date | 2022-03-07 02:49:21 UTC |
|---|
| HMDB ID | HMDB0004692 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | 12(R)-HPETE |
|---|
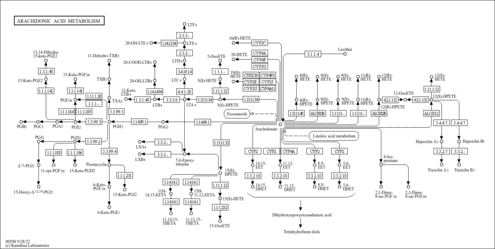
| Description | 12(R)-HPETE is a hydroperoxyeicosatetraenoic acid eicosanoid derived from arachidonic acid. The epidermal lipoxygenases 12R-LOX and eLOX3 act in sequence to convert arachidonic acid via 12(R)-HPETE to 12(R)-HETE and the corresponding epoxyalcohol, 8(R)-hydroxy-11(R),12(R)-epoxyeicosatrienoic acid. The epidermal lipoxygenases 12R-LOX and eLOX3 are the gene products of ALOX12B and ALOXE3. Mutations in ALOXE3 or ALOX12B have been found in families with autosomal-recessive congenital ichthyosis (ARCI). ARCI is a clinically and genetically heterogeneous group of severe hereditary keratinization disorders characterized by intense scaling of the whole integument, and differences in color and shape, often associated with erythema. Mutations in ALOXE3 and ALOX12B on chromosome 17p13, which code for two different epidermal lipoxygenases, were found in patients with ichthyosiform erythroderma. Genetic studies indicated that 12R-lipoxygenase (12R-LOX) or epidermal lipoxygenase-3 (eLOX3) was mutated in six families affected by non-bullous congenital ichthyosiform erythroderma (NCIE), one of the main clinical forms of ichthyosis. (PMID: 16116617 , 15629692 ). |
|---|
| Structure | CCCCC\C=C/C[C@@H](OO)\C=C\C=C/C\C=C/CCCC(O)=O InChI=1S/C20H32O4/c1-2-3-4-5-10-13-16-19(24-23)17-14-11-8-6-7-9-12-15-18-20(21)22/h7-11,13-14,17,19,23H,2-6,12,15-16,18H2,1H3,(H,21,22)/b9-7-,11-8-,13-10-,17-14+/t19-/m1/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (5Z,8Z,10E,14Z)-(12R)-12-Hydroperoxyeicosa-5,8,10,14-tetraenoic acid | ChEBI | | (5Z,8Z,10E,14Z)-(12R)-12-Hydroperoxyicosa-5,8,10,14-tetraenoic acid | ChEBI | | (5Z,8Z,10E,14Z)-(12R)-12-Hydroperoxyeicosa-5,8,10,14-tetraenoate | Generator | | (5Z,8Z,10E,14Z)-(12R)-12-Hydroperoxyicosa-5,8,10,14-tetraenoate | Generator | | 12R-HpETE | HMDB | | 12R-Hydroperoxy-5Z,8Z,10E,14Z-eicosatetraenoate | HMDB | | 12R-Hydroperoxy-5Z,8Z,10E,14Z-eicosatetraenoic acid | HMDB | | 12R-Hydroperoxyeicosatetraenoate | HMDB | | 12R-Hydroperoxyeicosatetraenoic acid | HMDB |
|
|---|
| Chemical Formula | C20H32O4 |
|---|
| Average Molecular Weight | 336.4657 |
|---|
| Monoisotopic Molecular Weight | 336.230059512 |
|---|
| IUPAC Name | (5Z,8Z,10E,12R,14Z)-12-hydroperoxyicosa-5,8,10,14-tetraenoic acid |
|---|
| Traditional Name | 12R-HpETE |
|---|
| CAS Registry Number | 126873-49-2 |
|---|
| SMILES | CCCCC\C=C/C[C@@H](OO)\C=C\C=C/C\C=C/CCCC(O)=O |
|---|
| InChI Identifier | InChI=1S/C20H32O4/c1-2-3-4-5-10-13-16-19(24-23)17-14-11-8-6-7-9-12-15-18-20(21)22/h7-11,13-14,17,19,23H,2-6,12,15-16,18H2,1H3,(H,21,22)/b9-7-,11-8-,13-10-,17-14+/t19-/m1/s1 |
|---|
| InChI Key | ZIOZYRSDNLNNNJ-ZYBDYUKJSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as hydroperoxyeicosatetraenoic acids. These are eicosanoic acids with an attached hydroperoxyl group and four CC double bonds. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Eicosanoids |
|---|
| Direct Parent | Hydroperoxyeicosatetraenoic acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Hydroperoxyeicosatetraenoic acid
- Long-chain fatty acid
- Hydroperoxy fatty acid
- Fatty acid
- Unsaturated fatty acid
- Allylic hydroperoxide
- Hydroperoxide
- Carboxylic acid derivative
- Alkyl hydroperoxide
- Carboxylic acid
- Peroxol
- Monocarboxylic acid or derivatives
- Carbonyl group
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxide
- Organic oxygen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | Not Available |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized | Show more...
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - 12(R)-HPETE GC-MS (Non-derivatized) - 70eV, Positive | splash10-0nml-8491000000-0127633b6ae0909f1672 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - 12(R)-HPETE GC-MS (1 TMS) - 70eV, Positive | splash10-05dr-9334000000-7ccf269dc3d82ad47d27 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - 12(R)-HPETE GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 10V, Positive-QTOF | splash10-014i-0119000000-19caf32dc0daf75104b3 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 20V, Positive-QTOF | splash10-00ou-5975000000-f4a0ca5c1a3a9fb036e3 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 40V, Positive-QTOF | splash10-052f-9640000000-496b0d1583282ff83466 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 10V, Negative-QTOF | splash10-000i-0119000000-f328ff1db02305a6bcd0 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 20V, Negative-QTOF | splash10-014r-1659000000-87c05812516016895a3b | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 40V, Negative-QTOF | splash10-0a4i-9740000000-43688cc7fb27bb45f0c9 | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 10V, Negative-QTOF | splash10-000i-0009000000-7847d679fdeeabe38573 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 20V, Negative-QTOF | splash10-0udr-0329000000-d4296bbebfcef533338a | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 40V, Negative-QTOF | splash10-056u-3690000000-c9e5a737326ebfdd3bf0 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 10V, Positive-QTOF | splash10-0f79-0249000000-702a285b6044fb920d29 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 20V, Positive-QTOF | splash10-0f79-7977000000-3cd8fc7d5c58fa9ef000 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 12(R)-HPETE 40V, Positive-QTOF | splash10-066u-9420000000-b47740fabc6ca56ad6c0 | 2021-09-24 | Wishart Lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | |
|---|
| Biospecimen Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | None |
|---|
| Associated OMIM IDs | None |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB023408 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 7827808 |
|---|
| KEGG Compound ID | C14812 |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 9548885 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 34145 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | Not Available |
|---|
| MarkerDB ID | Not Available |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Porter, Ned A.; Dussault, Patrick; Breyer, Robert A.; Kaplan, Jere; Morelli, Joseph. The resolution of racemic hydroperoxides: a chromatography-based separation of perketals derived from arachidonic, linoleic, and oleic acid hydroperoxides. Chemical Research in Toxicology (1990), 3(3), 236-43. |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Eckl KM, Krieg P, Kuster W, Traupe H, Andre F, Wittstruck N, Furstenberger G, Hennies HC: Mutation spectrum and functional analysis of epidermis-type lipoxygenases in patients with autosomal recessive congenital ichthyosis. Hum Mutat. 2005 Oct;26(4):351-61. [PubMed:16116617 ]
- Yu Z, Schneider C, Boeglin WE, Brash AR: Mutations associated with a congenital form of ichthyosis (NCIE) inactivate the epidermal lipoxygenases 12R-LOX and eLOX3. Biochim Biophys Acta. 2005 Jan 5;1686(3):238-47. [PubMed:15629692 ]
|
|---|