| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Expected but not Quantified |
|---|
| Creation Date | 2008-08-14 18:12:37 UTC |
|---|
| Update Date | 2022-03-07 02:49:34 UTC |
|---|
| HMDB ID | HMDB0006893 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | 3a,7a-Dihydroxy-5b-cholestane |
|---|
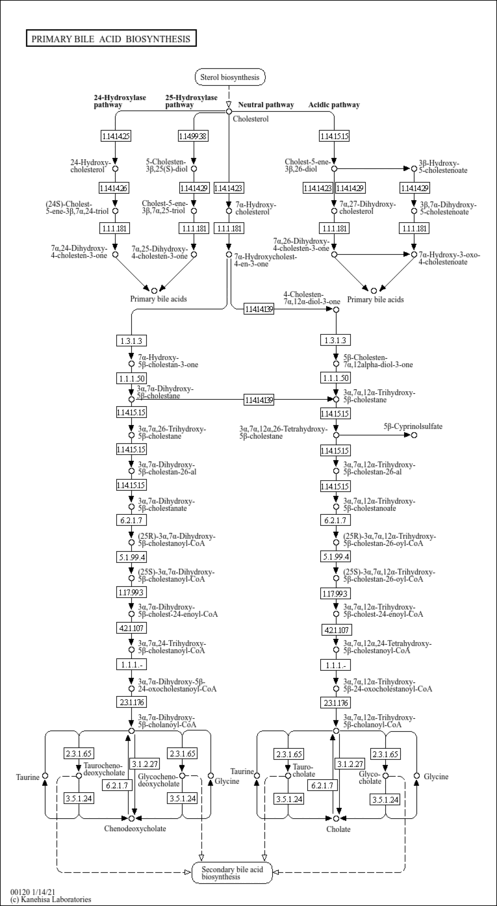
| Description | 3alpha,7alpha-Dihydroxy-5beta-cholestane is an intermediate in bile acid synthesis. Bile acids are steroid acids found predominantly in the bile of mammals. The distinction between different bile acids is minute, depending only on the presence or absence of hydroxyl groups on positions 3, 7, and 12. Bile acids are physiological detergents that facilitate excretion, absorption, and transport of fats and sterols in the intestine and liver. Bile acids are also steroidal amphipathic molecules derived from the catabolism of cholesterol. They modulate bile flow and lipid secretion, are essential for the absorption of dietary fats and vitamins, and have been implicated in the regulation of all the key enzymes involved in cholesterol homeostasis. Bile acids recirculate through the liver, bile ducts, small intestine and portal vein to form an enterohepatic circuit. They exist as anions at physiological pH and, consequently, require a carrier for transport across the membranes of the enterohepatic tissues. The unique detergent properties of bile acids are essential for the digestion and intestinal absorption of hydrophobic nutrients. Bile acids have potent toxic properties (e.g. membrane disruption) and there are a plethora of mechanisms to limit their accumulation in blood and tissues (PMID: 11316487 , 16037564 , 12576301 , 11907135 ). |
|---|
| Structure | [H][C@@]1(CC[C@@]2([H])[C@]3([H])[C@H](O)C[C@]4([H])C[C@H](O)CC[C@]4(C)[C@@]3([H])CC[C@]12C)[C@H](C)CCCC(C)C InChI=1S/C27H48O2/c1-17(2)7-6-8-18(3)21-9-10-22-25-23(12-14-27(21,22)5)26(4)13-11-20(28)15-19(26)16-24(25)29/h17-25,28-29H,6-16H2,1-5H3/t18-,19+,20-,21-,22+,23+,24-,25+,26+,27-/m1/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| 3alpha,7alpha-Dihydroxy-5beta-cholestane | ChEBI | | Dihydroxycoprostane | ChEBI | | 5beta-Cholestane-3alpha,7alpha-diol | Kegg | | 3Α,7α-dihydroxy-5β-cholestane | Generator | | 5b-Cholestane-3a,7a-diol | Generator | | 5Β-cholestane-3α,7α-diol | Generator | | 5alpha-Cholestane-3beta,7beta-diol | MeSH | | Dihydroxycoprostane, (3alpha,5alpha,7alpha)-isomer | MeSH | | 3,7-Dihydroxy-5-cholestane | MeSH | | Dihydroxycoprostane, (3beta,5alpha,7alpha)-isomer | MeSH | | 3,7-Dihydroxycholestane | MeSH |
|
|---|
| Chemical Formula | C27H48O2 |
|---|
| Average Molecular Weight | 404.6688 |
|---|
| Monoisotopic Molecular Weight | 404.36543078 |
|---|
| IUPAC Name | (1S,2S,5R,7S,9R,10R,11S,14R,15R)-2,15-dimethyl-14-[(2R)-6-methylheptan-2-yl]tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecane-5,9-diol |
|---|
| Traditional Name | 5β-cholestane-3α,7α-diol |
|---|
| CAS Registry Number | 3862-26-8 |
|---|
| SMILES | [H][C@@]1(CC[C@@]2([H])[C@]3([H])[C@H](O)C[C@]4([H])C[C@H](O)CC[C@]4(C)[C@@]3([H])CC[C@]12C)[C@H](C)CCCC(C)C |
|---|
| InChI Identifier | InChI=1S/C27H48O2/c1-17(2)7-6-8-18(3)21-9-10-22-25-23(12-14-27(21,22)5)26(4)13-11-20(28)15-19(26)16-24(25)29/h17-25,28-29H,6-16H2,1-5H3/t18-,19+,20-,21-,22+,23+,24-,25+,26+,27-/m1/s1 |
|---|
| InChI Key | APYVEUGLZHAHDJ-TVRYRFOISA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as cholesterols and derivatives. Cholesterols and derivatives are compounds containing a 3-hydroxylated cholestane core. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Cholestane steroids |
|---|
| Direct Parent | Cholesterols and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Cholesterol-skeleton
- Cholesterol
- 7-hydroxysteroid
- 3-alpha-hydroxysteroid
- Hydroxysteroid
- 3-hydroxysteroid
- Cyclic alcohol
- Secondary alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Organooxygen compound
- Alcohol
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. | 8.59 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 22.7482 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 2.55 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 47.7 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 3532.4 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 568.3 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 280.2 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 220.1 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 658.0 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 1121.8 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 964.3 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 99.2 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 1767.0 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 693.3 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 2169.7 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 574.2 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 580.4 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 220.7 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 487.9 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 8.4 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| 3a,7a-Dihydroxy-5b-cholestane,1TMS,isomer #1 | CC(C)CCC[C@@H](C)[C@H]1CC[C@H]2[C@H]3[C@H](CC[C@@]21C)[C@@]1(C)CC[C@@H](O)C[C@H]1C[C@H]3O[Si](C)(C)C | 3163.1 | Semi standard non polar | 33892256 | | 3a,7a-Dihydroxy-5b-cholestane,1TMS,isomer #2 | CC(C)CCC[C@@H](C)[C@H]1CC[C@H]2[C@H]3[C@H](CC[C@@]21C)[C@@]1(C)CC[C@@H](O[Si](C)(C)C)C[C@H]1C[C@H]3O | 3234.8 | Semi standard non polar | 33892256 | | 3a,7a-Dihydroxy-5b-cholestane,2TMS,isomer #1 | CC(C)CCC[C@@H](C)[C@H]1CC[C@H]2[C@H]3[C@H](CC[C@@]21C)[C@@]1(C)CC[C@@H](O[Si](C)(C)C)C[C@H]1C[C@H]3O[Si](C)(C)C | 3125.1 | Semi standard non polar | 33892256 | | 3a,7a-Dihydroxy-5b-cholestane,1TBDMS,isomer #1 | CC(C)CCC[C@@H](C)[C@H]1CC[C@H]2[C@H]3[C@H](CC[C@@]21C)[C@@]1(C)CC[C@@H](O)C[C@H]1C[C@H]3O[Si](C)(C)C(C)(C)C | 3408.9 | Semi standard non polar | 33892256 | | 3a,7a-Dihydroxy-5b-cholestane,1TBDMS,isomer #2 | CC(C)CCC[C@@H](C)[C@H]1CC[C@H]2[C@H]3[C@H](CC[C@@]21C)[C@@]1(C)CC[C@@H](O[Si](C)(C)C(C)(C)C)C[C@H]1C[C@H]3O | 3471.3 | Semi standard non polar | 33892256 | | 3a,7a-Dihydroxy-5b-cholestane,2TBDMS,isomer #1 | CC(C)CCC[C@@H](C)[C@H]1CC[C@H]2[C@H]3[C@H](CC[C@@]21C)[C@@]1(C)CC[C@@H](O[Si](C)(C)C(C)(C)C)C[C@H]1C[C@H]3O[Si](C)(C)C(C)(C)C | 3624.1 | Semi standard non polar | 33892256 |
|
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane GC-MS (Non-derivatized) - 70eV, Positive | splash10-06rl-1219000000-6af223a41b0a7a91af2c | 2017-09-01 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane GC-MS (2 TMS) - 70eV, Positive | splash10-001i-3220590000-375bc120297d486a3b68 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 10V, Positive-QTOF | splash10-052r-0009200000-df25d1509241d9f73f0a | 2016-08-03 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 20V, Positive-QTOF | splash10-05n0-2129100000-1bfe2b042bf2b32bbabd | 2016-08-03 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 40V, Positive-QTOF | splash10-0ab9-4129000000-cfef80f1eeab9ccd9236 | 2016-08-03 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 10V, Negative-QTOF | splash10-0udi-0003900000-9badc48971b46507f711 | 2016-08-03 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 20V, Negative-QTOF | splash10-0udr-0009800000-a236fdccb8989fe2c557 | 2016-08-03 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 40V, Negative-QTOF | splash10-00dr-1009000000-ac762387fbd63dd8fee4 | 2016-08-03 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 10V, Positive-QTOF | splash10-0a4i-0003900000-b8b020b5e69b09c721f9 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 20V, Positive-QTOF | splash10-0ab9-9013100000-34ed7551b67a9fc921bc | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 40V, Positive-QTOF | splash10-0a4m-9430000000-dd74d7aaa66a164f94b8 | 2021-09-22 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 10V, Negative-QTOF | splash10-0udi-0000900000-82ba826e352c471b5c11 | 2021-09-23 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 20V, Negative-QTOF | splash10-0udi-0000900000-3db0959145384e45d712 | 2021-09-23 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 3a,7a-Dihydroxy-5b-cholestane 40V, Negative-QTOF | splash10-0udi-0000900000-60f92d1adbf02fcd18e9 | 2021-09-23 | Wishart Lab | View Spectrum |
|
|---|